Detection of SARS-CoV-2 in Nigeria: laboratory preparedness and report of index case
Ayorinde Babatunde James, Olumuyiwa Babalola Salu, Joseph Ojonugwa Shaibu, Chika Onwuamah, Bamidele Paul Mutiu, Olufemi Amoo, Mercy Remilekun Orenolu, Azuka Okwuraiwe, Nnaemeka Ndodo, Babatunde Akeem Saka, Roosevelt Amaobichukwu Anyanwu, Munir Akinwale Bankole, Fehintola Ige, Mariam Abiodun Abdullah, Ismail Adeshina Abdus-Salam, Rosemary Ajuma Audu, Akin Osibogun, Akin Abayomi, Chikwe Ihekweazu, Babatunde Lawal Salako, Sunday Aremu Omilabu
Corresponding author: Sunday Aremu Omilabu, Centre for Human and Zoonotic Virology (CHAZVY), Central Research Laboratory, College of Medicine, University of Lagos, Idi-Araba, Lagos, Nigeria 
Received: 12 Jan 2021 - Accepted: 05 May 2021 - Published: 21 May 2021
Domain: Virology,Infectious diseases epidemiology
Keywords: Coronavirus, SARS-CoV-2, MERS-CoV, Betacoronavirus, qPCR, sequencing, Nigeria
©Ayorinde Babatunde James et al. PAMJ-One Health (ISSN: 2707-2800). This is an Open Access article distributed under the terms of the Creative Commons Attribution International 4.0 License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Cite this article: Ayorinde Babatunde James et al. Detection of SARS-CoV-2 in Nigeria: laboratory preparedness and report of index case. PAMJ-One Health. 2021;5:9. [doi: 10.11604/pamj-oh.2021.5.9.27817]
Available online at: https://www.one-health.panafrican-med-journal.com/content/article/5/9/full
Detection of SARS-CoV-2 in Nigeria: laboratory preparedness and report of index case
Ayorinde Babatunde James1, ![]() Olumuyiwa Babalola Salu2, Joseph Ojonugwa Shaibu3,
Olumuyiwa Babalola Salu2, Joseph Ojonugwa Shaibu3, ![]() Chika Onwuamah3, Bamidele Paul Mutiu4,
Chika Onwuamah3, Bamidele Paul Mutiu4, ![]() Olufemi Amoo3, Mercy Remilekun Orenolu2,
Olufemi Amoo3, Mercy Remilekun Orenolu2, ![]() Azuka Okwuraiwe3, Nnaemeka Ndodo5,
Azuka Okwuraiwe3, Nnaemeka Ndodo5, ![]() Babatunde Akeem Saka6, Roosevelt Amaobichukwu Anyanwu2, Munir Akinwale Bankole4, Fehintola Ige3, Mariam Abiodun Abdullah2, Ismail Adeshina Abdus-Salam7,
Babatunde Akeem Saka6, Roosevelt Amaobichukwu Anyanwu2, Munir Akinwale Bankole4, Fehintola Ige3, Mariam Abiodun Abdullah2, Ismail Adeshina Abdus-Salam7, ![]() Rosemary Ajuma Audu3,
Rosemary Ajuma Audu3, ![]() Akin Osibogun8, Akin Abayomi9, Chikwe Ihekweazu5,
Akin Osibogun8, Akin Abayomi9, Chikwe Ihekweazu5, ![]() Babatunde Lawal Salako10,
Babatunde Lawal Salako10, ![]() Sunday Aremu Omilabu2,3
Sunday Aremu Omilabu2,3
&Corresponding author
Introduction: heightened surveillance of the severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) 2019 outbreak in Wuhan, China activated several collaborative laboratory drills on its detection by health institutions in Nigeria. We hereby detail the laboratory detection and confirmation of the novel SARS-CoV-2 virus of the index case imported into Nigeria.
Methods: viral RNA from sputum sample of the index case was tested for SARS-CoV-2 envelope (E) and RNA dependent RNA polymerase (RdRp) genes using quantitative reverse transcription-polymerase chain reaction (qRT-PCR). Genetic analysis using a partial sequence of RdRp gene was used for laboratory confirmation, evolutionary inferences, and relatedness to previous strains of betacoronaviruses in Nigeria.
Results: both qRT-PCR for the E and RdRp genes indicated the presence of the SARS-CoV-2 RNA by showing cycle threshold (ct) values of 20.27 and 25.26 respectively. On successful completion of the sequencing run, data aligned with >98% similarity of the 2019 circulating strain of Coronavirus SARS-CoV-2. Evolutionary algorithms revealed that the sequence (RdRp gene) of the outbreak circulating strains were more conserved unlike previous coronavirus outbreaks. Data showed that some coronaviruses belonging to the BetaCoV genus had earlier been reported in bats and camels in Nigeria indicating the possibility of sero-protection in exposed populations.
Conclusion: pre-laboratory drills prompted early diagnosis and confirmation of SARS-CoV-2 index case in Nigeria initiating rapid case-definition, contact tracing, and successful epidemiological management of the outbreak. Rapid sequencing of the viral genome during strategic points of the outbreak curve and making libraries of new variants for improved vaccine development are recommended.
Coronaviruses (CoVs) belongs to the family of a large group of non-segmented positive single-stranded RNA (27-32Kb genome length) viruses. These group of viruses cause significant diseases in humans such as severe acute respiratory syndrome (SARS) and other infections in an increasingly large variety of mammals and birds [1]. They are enveloped viruses sized 80 - 220 nm in diameter with club-shaped spike projections emanating from the surface of the virion. These corona- or crown-like spikes are a defining feature of the virion and give them the appearance of a solar corona [2]. They are members of the subfamily Coronavirinae in the family Coronaviridae and the order Nidovirales. This subfamily divides into 4 genera - alphacoronavirus, betacoronavirus, gammacoronavirus, and deltacoronavirus [3]. Alpha and beta-coronaviruses infects mainly mammals while delta and gamma-coronaviruses are commonly known to infect birds [4]. In humans, four coronaviruses (HCoV-229E, OC43, NL63, and HKU1) are endemic causing common cold. In addition to these four endemic coronaviruses, two Beta coronaviruses namely severe acute respiratory syndrome (SARS-CoV) and the Middle East Respiratory Syndrome (MERS-CoV) were responsible for outbreaks of viral pneumonia in 2003 and 2012 respectively [5].
In December 2019, a novel coronavirus was identified amongst patients with unexplained cause of pneumonia in Wuhan, China [6,7]. The World Health Organization (WHO) referred to the virus disease as COVID-19. Due to global travels, it took no time to become one of the worst human pandemics in the 21st century. By the 30th of January 2020, WHO declared the outbreak a Public Health Emergency of International Concern (PHEIC). On 11th March 2020, the WHO Director General characterized COVID-19 as a pandemic [8]. Prior to the declaration of the 2019 severe acute respiratory disease (SARS-CoV-2) as an epidemic in Wuhan Province, China; the Nigerian Centre for Disease Control (NCDC), Nigerian Institute of Medical Research (NIMR), Lagos State Ministry of Health, and the Centre for Human and Zoonotic Virology (CHAZVY) of the College of Medicine-University of Lagos had commenced laboratory drills on qRT-PCR protocol testing, optimization using recombinant control specimens, and sequencing of generated PCR products. Past occurrences of highly pathogenic Avian/H5N1 in 2006 [9], Swine-flu (H1N1/09) in 2009 [10], and Ebola viral disease importation into Nigeria in 2014 [11] served as important lessons in strengthening laboratory infrastructure, preparedness, and mitigation efforts in outbreak situations. These situations have led to the establishment of the Avian influenza project in Nigeria and the Viral haemorrhagic fever task force leading to the establishment of Molecular testing laboratories and sentinel surveillance sites in various geo-political zones of Nigeria.
Nigeria confirmed its first case of COVID-19 on the 27th February 2020 at the Infectious Disease Hospital, Yaba, Lagos. The index case was a 44-year-old Italian male expatriate who returned from a family visit in Italy. Replicate samples were shared with NCDC network laboratories for confirmation and genetic analysis. Hence a case definition for testing including all symptomatic individuals with travel history to China or COVID-19 endemic countries was established. In an effort to rapidly diagnose the suspected case, confirm its strain using genetic analysis, and investigate if Nigeria had reported any other case of Beta-coronavirus prior to the COVID-19 pandemic, we present our account of the laboratory detection and confirmation of the novel SARS-CoV-2 virus of the index case imported into Nigeria using quantitative reverse transcription-polymerase chain reaction (qRT-PCR) and genetic analysis.
Sample: sputum sample was collected in a viral transport medium (VTM) and sent to the Centre for Human and Zoonotic Virology (CHAZVY), College of Medicine-University of Lagos (CMUL)/Lagos University Teaching Hospital (LUTH) for quantitative real time PCR detection using the diagnostic algorithm for the detection of coronaviruses by the Nigeria Centre for Disease Control (NCDC) and World Health Organization (WHO) [12].
Ethical considerations: this detection and confirmation investigations were performed as part of the COVID-19 public health response in Nigeria. Protocol was reviewed and approved (IRB/20/101) by the Institutional Review Board of the Nigerian Institute of Medical Research and was considered an analysis of public health service data as documented by Otto, Holodniy [13]. All data were completely anonymised before analysis.
Sample processing and nucleic acid extraction: sputum sample was processed for inactivation in a class-3 glove box at the Lagos State Biosafety and Biobank Laboratory by adding 140 μL of sample to 560 μL of viral lysis buffer of Qiagen Viral RNA Extraction Kit in a label tube (Qiagen, MD). The mixture was placed on a heating block at 56°C for 20 mins with periodic pulse vortexing. Subsequently, viral RNA extraction (QIAamp Viral RNA mini kit, Qiagen USA) of the sputum sample was carried out in a class 2 biosafety cabinet. All procedures were carried out according to World Health Organization´s (WHO) general procedures for inactivation of viral pathogens in a BSL3 facility [14,15].
Quantitative polymerase chain reaction (qPCR) virus detection: viral RNA from the sputum sample was assayed for COVID-19 at CHAZVY and aliquots of viral RNA were shared amongst laboratories in the NCDC network and the Nigerian Institute of Medical Research. An initial RT-PCR assay was setup using the kit supplied by the NCDC with oligonucleotides from TIB MOLBIOL LightMix® Modular SARS-CoV (Berlin) oligonucleotides targeting envelope (E) gene and RNA dependent RNA polymerase (RdRp) gene (Table 1). The reaction was carried out using SuperScript™ III Platinum™ One-Step qRT-PCR Kit (ThermoFisher Scientific, USA) in a 25 μL containing 1X reaction mix, 0.8 mM MgSO4, 0.04 mg/mL BSA, 0.4 μM each of forward and reverse primers, 0.2 μM probe, 1 μL of SSIII/Taq Enzyme mix, and 5 μL purified RNA. For the RdRp assay, forward primer concentration was increased to 0.6 μM. Cycling was done at 55°C for 10 minutes, 95 °C for 3 minutes hold, two step 45X repeat cycles of 94 °C for 15 seconds and 58 °C for 30 seconds. Fluorescent data capture was acquired at FAM and Cy5 channels during each second cycling step. Subsequently, One-step RT-qPCR was carried out for the detection of SARS-CoV-2 virus using assay kit designed by BGI, Shenzhen, China at NIMR. The reaction contained 18.5 μL of SARS-CoV-2 nucleic acid mix, 1.5 μL of enzyme mix and 10 μL of RNA in a total volume of 30 μL. Real-time PCR cycling was performed on QuantStudio-3 Applied Biosystems Real-Time PCR machine as follows: 50°C for 20 mins, 95°C for 10 mins, then 40 cycles of 15s at 95°C and 30s at 60°C.
Amplicon sequencing of viral RNA Dependent RNA polymerase (RdRp): for the purpose of sanger sequencing at the Nigerian Institute of Medical Research, primers (Table 1) reported by Pauly, Pir [16] were used in a 25 μL reaction consisting of 1X SCRIPT RT-PCR Reaction Mix (Jena Bioscience, Germany), 1mM forward primer, 1mM reverse primer, 1 μL SCRIPT RT-PCR Enzyme Mix, and 5 μL of RNA template Cycling condition was done at 50°C for 40 min, initial denaturation at 95°C for 5 min, 95°C for 10 sec, 57°C for 20 sec, and 72°C for 1 min for 40 cycles with a final extension at 72°C for 5 min. PCR products were separated on 2% Tris-acetate-EDTA (TAE) agarose gel electrophoresis for 30 min at 120 V. Observed PCR bands were excised and purified using ZymocleanTM Gel DNA Recovery Kit (Zymo Research, USA). Prior to sequencing, gel purified products were quantified using Qubit 4 fluorometer (ThermoFisher Scientific, CA). Sequencing was performed using the BigDye Terminator kit (Applied Biosystems, Foster City, CA), run on an ABI 3130-xl and SeqStudio Genetic Analyzers (Applied Biosystems). Partial sequencing of SARS-CoV-2 was obtained using nested primers targeting the conserved RdRp gene (Table 1).
Phylogenetic analysis: obtained sequence was searched against sequences in the GenBank using the blastn search tool. Sequences related to the current outbreak, alpha, beta, gamma, and delta coronaviruses were retrieved using the research and efetch Linux command tools of BLAST 2.6.0+. Over one hundred retrieved sequences were aligned using the muscle command [17], viewed and edited with belvu tools [18]. The best substitution model required for constructing a Molecular likelihood (ML) tree was determined to be General Time Reversible (GTR) using a discrete Gamma distribution (+G) according to the Bayesian information criterion (BIC) and/or Akaike information criterion (AIC) values. This analysis was done using MegaX software and it involved 82 nucleotide sequences and a bootstrap test including 1,000 replicates was performed for the ML tree [19].
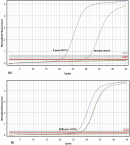
qPCR virus detection: using the kits provided by the Nigeria Centre for Disease Control (NCDC) targeting the E and RdRp gene segments [20], a cycle threshold (Ct) values of 20.27 and 25.26 was observed for the E and RdRp genes of RNA extract of the sputum sample at CHAZVY as shown in Figure 1. A cycle threshold (Ct) value of 28.2 was also observed on the subsequent run using the QuantStudio-3 Real Time PCR system at NIMR. Further analysis at NIMR using the BGI-China kits also corroborated the qPCR result by showing a Ct value of 28 for the orf1ab gene segment. The evidences from these runs indicates the presence of an active SARS-CoV-2 infection.
Sanger sequencing: the short sequence targeting the RdRp gene generated during the course of this investigation further established the diagnosis of COVID-19. A clean chromatogram (Figure 2) removes chances of ambiguity and poor sequence base calling. Sequence extracted from this information was used for phylogenetic analysis to investigate the evolutionary relationship between the obtained raw sequence and existing COVID-19 outbreak sequences including Beta, Alpha, and Gamma CoV sequences.
Sequence alignment and phylogeny: sequence alignment data of the Lagos index case with other COVID-19 sequences generated from other parts of the world revealed a high level of conservation in the RdRp region. This seems to be a deviant from other BetaCoV species that are highly variable in the same RdRp region because RNA viruses mutation rate is very high enabling adaptability and stability [21]. Phylogenetic analysis as shown in Figure 3 revealed that the obtained sequence of the Lagos index case clustered (95% bootstrapping) with the current COVID-19 outbreak sequences which are part of the SARS related CoV of the Beta-CoV genus. All the sequences in this outbreak were found to cluster with earlier reported sequence data from isolates in SARS coronavirus Shanghai (accession no. AY463060), civet mammal in south China (AY304488), and bat coronavirus (DQ412042) in Wuhan China [22] submitted to the Genbank between 2003 and 2004.
A BetaCoV-1 coronavirus sequence data (HQ166910) isolated from bats (Hipposideros commersoni) in Zaria, Nigeria in 2008 and MERS-CoV (MG923476) from camels in Nigeria were submitted in 2016. This information indicates that we had circulating strains of Beta-CoV genus in Nigeria and not the novel SARS-CoV-2 coronavirus. Furthermore, the phylogenetic tree revealed that some species of the same BetaCoV genus had been isolated from other animals such as rats, mice, dogs, horse, cow, deer and pigs in many parts of the world.
Nucleic acid detection methods such as polymerase chain reaction (PCR), array hybridizations, and sequencing have gained revolutionary prominence as the gold standard for virus detection and confirmation due to its speed, sensitivity, specificity, and robustness in outbreak situations [23,24]. The Nigerian experience with the scale up of laboratory facilities and the increased training of laboratory personnel to conduct molecular testing has been of immense benefit to the country in stemming outbreaks that would have hitherto had catastrophic effects in the most populous black nation in the world. In our report, quantitative RT-PCR technique proved effective in the rapid diagnosis of COVID-19 with a turnaround time as low as six hours which assisted in establishing a case definition for the outbreak and instant contact tracing. For the purpose of genetic characterization of the viral isolate, the genetic analyzer still remains a good source in providing nucleic acid sequence information within 24 hours with less cumbersome bioinformatic analysis experienced during analysis of next generation (NGS) data. Nonetheless, NGS platforms are still recommended for whole genome sequencing in order to provide greater coverage and improved sequencing depths [25]. The short sequence generated during the course of our investigation further established the diagnosis of COVID-19 in Nigeria. The African Centre of Excellence for the Genomics of Infectious Diseases (ACEGID) at Redeemer´s University, Ede, Nigeria (RUN) went on further to provide the whole genome sequence of SARS-CoV-2 after diagnosis of COVID-19 had been made in the index patient in Nigeria [26]. In addition, Data mined by Awoyelu, Oladipo [27] from GISAID using whole genome sequences showed a 100% similarity implicating a common origin of evolution.
Sequence data from the RdRp region are known to be commonly used tool for the detection of coronavirus, identification, and molecular epidemiology classification [28, 29]. Sequence data identified the Nigerian isolate as a Sarbecovirus of the Betacoronavirus genus and it was named SARS-CoV-2/human/NGA/NG01/2020 and deposited with the National Center for Biotechnology Institute´s (NCBI) GenBank. Our phylogenetic analysis implicated isolates from bats as the major feeder of outbreaks in previous Betacoronavirus outbreaks and the ongoing COVID-19 pandemic. This was affirmed by earlier suggestions and reports by Drexler, Corman [30] and Hu, Ge [22] suggesting that severe acute respiratory syndrome CoV (SARS-CoV) and the Middle East respiratory syndrome CoV (MERS-CoV) originated in bats.
Despite high levels of genetic mutations observed in RNA viruses, we observed that the partial sequence generated is relatively conserved across different reported sequences from other countries involved in this pandemic [1]. More so, the RdRp region of coronaviruses show less variability than the spike region [31]. This may indicate that the virus is slowly mutating and may have a seemingly stable genome which is good news for effective and consistent vaccine. Nonetheless, healthcare authorities should continue to monitor the evolution of the viral genome during the outbreak incase another variant arises [32]. Our report also revealed that SARS-Like coronaviruses (SL-CoV) were isolated from bats and camels in Northern part of Nigeria. The Zaria bat coronavirus (Genbank Accession No. HQ166910.1) had 95% similarity with SARS-CoV-2 and the outbreak feeder strains from China. Similar isolates were found in Ghana and Kenya which when compared with Asian and European SL-CoVs showed phylogenetic distance to SARS-CoVs [33-35].
Based on our findings, we would like to recommend that more rapid and cost-effective nucleic acid point of care tests be developed and deployed for field testing; engage in continuous sequencing of the viral genome during contact tracing and in animals during strategic points of the outbreak curve; make cDNA libraries of new variants of the virus for possible vaccine development strategies.
Funding: this research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors. All materials used were provided by various institutions involved in this report.
The authors declare no competing interests.
AA, IC, AA, OA, BSL, and OSA led outbreak management. OA, AIA, BMA, SBA supervised the field outbreak management. ARA, OSA, CK, and SOB coordinated the laboratory response in the outbreak. OA, NN, IF, AMA, ARA, AR, AO, OMR, JOS, and JAB conducted the laboratory investigations. JAB, JOS, ARA, and SBL analyzed the data. Figures and tables were generated by JAB and SOB. JAB and ARA drafted the report and reviewed by all the authors. All the authors have read and agreed to the final manuscript.
We acknowledge all the staff of collaborating institutions for their cooperation and support to ensure the success of this project.
Table 1: primers and probes used for detection and sequencing of coronaviruses
Figure 1: qRT-PCR Cycle Threshold (Ct) Values for the E gene (A) and the RdRp gene (B) on the Magnetic Induction Cycler (MIC PCR 48)
Figure 2: raw sequence chromatogram of COVID-19 partial RdRp gene segment. Trace files were generated from orf1ab amplicon sequencing using PCR primers reported by Pauly, Pir
Figure 3: phylogenetic analysis of the index case with respect to other sequences of coronaviruses. The Nigerian Index case's sequence is highlighted in a pink box (SARS-CoV-2/Human/LAG/NG01/02-2020) while the Chinese cases are shown in red font. The SARS-CoV-2 reference strain is highlighted in red font and denoted MN988668.1/2019-nCoV WHU01. Previously reported isolated BetaCoV-1 sequences in Nigeria are highlighted in blue boxes
- Denison MR, Graham RL, Donaldson EF, Eckerle LD, Baric RS. Coronaviruses: an RNA proofreading machine regulates replication fidelity and diversity. RNA Biol. 2011;8(2):270-279. PubMed | Google Scholar
- Fehr AR, Perlman S. Coronaviruses: an overview of their replication and pathogenesis. Methods Mol Biol. 2015;1282:1-23. PubMed | Google Scholar
- Ludwig S, Zarbock A. Coronaviruses and SARS-CoV-2. Anesth Analg. 2020 Jul;131(1):93-96. PubMed | Google Scholar
- Miłek J, Blicharz-Domańska K. Coronaviruses in avian species - review with focus on epidemiology and diagnosis in wild birds. J Vet Res. 2018 Dec 10;62(3):249-255. PubMed | Google Scholar
- Corman VM, Muth D, Niemeyer D, Drosten C. Hosts and sources of endemic human coronaviruses. Adv Virus Res. 2018;100:163-188. PubMed | Google Scholar
- Zhu N, Zhang D, Wang W, Li X, Yang B, Song J et al. A Novel coronavirus from patients with pneumonia in China, 2019. N Engl J Med. 2020;382(8):727-733. PubMed | Google Scholar
- WHO. Laboratory testing for 2019 novel coronavirus (2019-nCoV) in suspected human cases, Interim guidance, 2 March 2020. World Health Organization; 2020. Accessed 24 March 2020.
- WHO. Situation report-51 situation in numbers total and new cases in last 24 hours. World Health Organization; 2020. Accessed 21 March 2020.
- Perry BD, Randolph TF, Bett BK, Grace D, Globig A, Fadigba M et al. Independent impact assessment of the World Bank-funded Nigeria Avian Influenza Control and Human Pandemic Preparedness and Response Project (NAICP). 2011. International Livestock Research Institute, Nigeria. PubMed | Google Scholar
- Meseko CA, Odaibo GN, Olaleye DO. Detection and isolation of 2009 pandemic influenza A/H1N1 virus in commercial piggery, Lagos Nigeria. Vet. Microbiol. 2014;168(1):197-201. PubMed | Google Scholar
- Salu OB, James AB, Oke BO, Orenolu MR, Anyanwu RA, Abdullah MA et al. Biosafety level-2 laboratory diagnosis of Zaire Ebola virus disease imported from Liberia to Nigeria. Afr J Lab Med. 2016;5(1):1-5. PubMed | Google Scholar
- WHO. Laboratory testing for 2019 novel coronavirus (2019-nCoV) in suspected human cases, Interim guidance 17 January 2020. World Health Organization; 17 January 2020. Accessed 24 March 2020.
- Otto JL, Holodniy M, DeFraites RF. Public health practice is not research. Am J Public Health. 2014;104(4):596-602. PubMed | Google Scholar
- Hartman AL, Cole KS, Homer LC. Verification of inactivation methods for removal of biological materials from a biosafety level 3 select agent facility. Appl. Biosaf. 2012;17(2):70-75. Google Scholar
- WHO. Laboratory biosafety guidance related to coronavirus disease 2019 (COVID-19): interim guidance, 12 February 2020. Accessed 24 March 2020.
- Pauly M, Pir JB, Loesch C, Sausy A, Snoeck CJ, Hübschen JM et al. Novel Alphacoronaviruses and paramyxoviruses cocirculate with type 1 and severe acute respiratory system (SARS)-related betacoronaviruses in synanthropic bats of Luxembourg Appl Environ. Microbiol. 2017;83(18):1-14. PubMed | Google Scholar
- Edgar RC. Muscle: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32(5):1792-1797. PubMed | Google Scholar
- Sonnhammer ELL, Hollich V. Scoredist: a simple and robust protein sequence distance estimator. BMC Bioinformatics. 2005 Apr 27;6:108. PubMed | Google Scholar
- Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol. 2018;35(6):1547-1549. PubMed | Google Scholar
- Corman VM, Landt O, Kaiser M, Molenkamp R, Meijer A, Chu DKW et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro surveill. 2020;25(3):1-8. PubMed | Google Scholar
- Pachetti M, Marini B, Benedetti F, Giudici F, Mauro E, Storici P et al. Emerging SARS-CoV-2 mutation hot spots include a novel RNA-dependent-RNA polymerase variant. J Transl Med. 2020;18(1):1-9. PubMed | Google Scholar
- Hu B, Ge X, Wang LF, Shi Z. Bat origin of human coronaviruses. Virol J. 2015;12(1):221-231. PubMed | Google Scholar
- Shen M, Zhou Y, Ye J, Abdullah Al-maskri AA, Kang Y, Zeng S et al. Recent advances and perspectives of nucleic acid detection for coronavirus. J Pharm. Anal. 2020;10(2):97-101. PubMed | Google Scholar
- Storch GA. Diagnostic Virology. Clin Infect Dis. 2000;31(3):739-751. PubMed | Google Scholar
- Hagemann IS. Overview of technical aspects and Chemistries of Next-Generation Sequencing. In Clinical Genomics. 2015. Elsevier. 2015;3-19. Google Scholar
- Olaniyi P, Ihekweazu C, Happi C, Omilabu S, Salako B, Abayomi A. First African SARS-CoV-2 genome sequence from Nigerian COVID-19 case. Genome Reports. 1 March 2020.
- Awoyelu EH, Oladipo EK, Adetuyi BO, Senbadejo TY, Oyawoye OM, Oloke JK. Phyloevolutionary analysis of SARS-CoV-2 in Nigeria. New microbes new infect. 2020 Dec;101(12):1261-1269. PubMed | Google Scholar
- Wilkinson DA, Joffrin L, Lebarbenchon C, Mavingui P. Analysis of partial sequences of the RNA-dependent RNA polymerase gene as a tool for genus and subgenus classification of coronaviruses. J Gen Virol. 2020;101:1261-1269. PubMed | Google Scholar
- Lehmann KC, Gulyaeva A, Zevenhoven-Dobbe JC, Janssen GM, Ruben M et al. Discovery of an essential nucleotidylating activity associated with a newly delineated conserved domain in the RNA polymerase-containing protein of all nidoviruses. Nucleic Acids Res. 2015;43(17):8416-8434. PubMed | Google Scholar
- Drexler JF, Corman VM, Drosten C. Ecology, evolution and classification of bat coronaviruses in the aftermath of SARS. Antiviral Res. 2014;101:45-56. PubMed | Google Scholar
- Zhang WF, Stephen P, Thériault JF, Wang R, Lin SX. Novel Coronavirus Polymerase and Nucleotidyl-Transferase Structures: Potential to Target New Outbreaks. J. Phys. Chem. Lett. 2020;11(11):4430-4435. PubMed | Google Scholar
- Khailany RA, Safdar M, Ozaslan M. Genomic characterization of a novel SARS-CoV-2. Gene Rep. 2020 Jun;19:100682. PubMed | Google Scholar
- Pfefferle S, Oppong S, Drexler JF, Gloza-Rausch F, Ipsen A, Seebens A et al. Distant relatives of severe acute respiratory syndrome coronavirus and close relatives of human coronavirus 229E in bats, Ghana. Emerg. Infect. Dis. 2009;15(9):1377-1384. PubMed | Google Scholar
- Tong S, Conrardy C, Ruone S, Kuzmin IV, Guo X, Tao Y et al. Detection of novel SARS-like and other coronaviruses in bats from Kenya. Emerg. Infect. Dis. 2009;15(3):482-485. PubMed | Google Scholar
- Quan P-L, Firth C, Street C, Henriquez JA, Petrosov A, Tashmukhamedova A et al. Identification of a severe acute respiratory syndrome coronavirus-like virus in a leaf-nosed bat in Nigeria. mBio. 2010;1(4):e00208-00210. PubMed | Google Scholar
Search
This article authors
On Pubmed
- Ayorinde Babatunde James
- Olumuyiwa Babalola Salu
- Joseph Ojonugwa Shaibu
- Chika Onwuamah
- Bamidele Paul Mutiu
- Olufemi Amoo
- Mercy Remilekun Orenolu
- Azuka Okwuraiwe
- Nnaemeka Ndodo
- Babatunde Akeem Saka
- Roosevelt Amaobichukwu Anyanwu
- Munir Akinwale Bankole
- Fehintola Ige
- Mariam Abiodun Abdullah
- Ismail Adeshina Abdus-Salam
- Rosemary Ajuma Audu
- Akin Osibogun
- Akin Abayomi
- Chikwe Ihekweazu
- Babatunde Lawal Salako
- Sunday Aremu Omilabu
On Google Scholar
- Ayorinde Babatunde James
- Olumuyiwa Babalola Salu
- Joseph Ojonugwa Shaibu
- Chika Onwuamah
- Bamidele Paul Mutiu
- Olufemi Amoo
- Mercy Remilekun Orenolu
- Azuka Okwuraiwe
- Nnaemeka Ndodo
- Babatunde Akeem Saka
- Roosevelt Amaobichukwu Anyanwu
- Munir Akinwale Bankole
- Fehintola Ige
- Mariam Abiodun Abdullah
- Ismail Adeshina Abdus-Salam
- Rosemary Ajuma Audu
- Akin Osibogun
- Akin Abayomi
- Chikwe Ihekweazu
- Babatunde Lawal Salako
- Sunday Aremu Omilabu
Citation [Download]
Navigate this article
Similar articles in
Key words
Tables and figures
 Figure 3: phylogenetic analysis of the index case with respect to other sequences of coronaviruses. The Nigerian Index case's sequence is highlighted in a pink box (SARS-CoV-2/Human/LAG/NG01/02-2020) while the Chinese cases are shown in red font. The SARS-CoV-2 reference strain is highlighted in red font and denoted MN988668.1/2019-nCoV WHU01. Previously reported isolated BetaCoV-1 sequences in Nigeria are highlighted in blue boxes
Figure 3: phylogenetic analysis of the index case with respect to other sequences of coronaviruses. The Nigerian Index case's sequence is highlighted in a pink box (SARS-CoV-2/Human/LAG/NG01/02-2020) while the Chinese cases are shown in red font. The SARS-CoV-2 reference strain is highlighted in red font and denoted MN988668.1/2019-nCoV WHU01. Previously reported isolated BetaCoV-1 sequences in Nigeria are highlighted in blue boxes







