Detection of ESBLs and NDM-1 genes among urinary Escherichia coli and Klebsiella pneumoniae from healthy students in Niger Delta University, Amassoma, Bayelsa State, Nigeria
Adebola Onanuga, Darlington Deboh Eboh, Babatunde Odetoyin, Ocholi Jonathan Adamu
Corresponding author: Adebola Onanuga, Department of Pharmaceutical Microbiology and Biotechnology Faculty of Pharmacy, Niger Delta University, Wilberforce Island Bayelsa State, Nigeria 
Received: 18 May 2020 - Accepted: 02 Jun 2020 - Published: 17 Jun 2020
Domain: Bacteriology,Infectious diseases epidemiology,Public health
Keywords: NDM-1, ESBLs, ASB, MDR, healthy individuals, community, Nigeria
©Adebola Onanuga et al. PAMJ-One Health (ISSN: 2707-2800). This is an Open Access article distributed under the terms of the Creative Commons Attribution International 4.0 License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Cite this article: Adebola Onanuga et al. Detection of ESBLs and NDM-1 genes among urinary Escherichia coli and Klebsiella pneumoniae from healthy students in Niger Delta University, Amassoma, Bayelsa State, Nigeria. PAMJ-One Health. 2020;2:12. [doi: 10.11604/pamj-oh.2020.2.12.23588]
Available online at: https://www.one-health.panafrican-med-journal.com/content/article/2/12/full
Research 
Detection of ESBLs and NDM-1 genes among urinary Escherichia coli and Klebsiella pneumoniae from healthy students in Niger Delta University, Amassoma, Bayelsa State, Nigeria
Detection of ESBLs and NDM-1 genes among urinary Escherichia coli and Klebsiella pneumoniae from healthy students in Niger Delta University, Amassoma, Bayelsa State, Nigeria
Adebola Onanuga1,2,&, Darlington Deboh Eboh1, Babatunde Odetoyin3, Ocholi Jonathan Adamu2
&Corresponding author
Introduction: increasing emergence of multidrug-resistant uropathogens among healthy individuals is a serious public health problem capable of causing difficult-to-treat urinary tract infections (UTIs) with limited treatment options. This study therefore, determined the prevalence of asymptomatic bacteriuria (ASB) and molecular characteristics of antibiotic-resistant Escherichia coli and Klebsiella pneumoniae isolated from healthy students of Niger Delta University, Amassoma, Nigeria.
Methods: mid-stream urine samples were collected from 303 healthy students and cultured for significant bacteriuria. Escherichia coli and K. pneumoniae were isolated and identified by conventional methods and Polymerase Chain Reaction (PCR). Antimicrobial susceptibility patterns of the isolates were determined by the disc diffusion technique. The isolates were screened for Extended-Spectrum Beta-Lactamases (ESBLs) production by combined disc method, ESBLs and carbapenemases genes by PCR.
Results: the prevalence of ASB was 21.1%, with a significantly higher proportion among the females (P < 0.0001). Escherichia coli and K. pneumoniae isolates were recovered in equal proportion (n=21; 6.9%), with E. coli significantly isolated from the females (P= 0.002). The isolates exhibited 57-95% resistance to cefotaxime, ceftazidime, and co-trimoxazole, and 10-24% resistance to levofloxacin and imipenem. Multidrug resistance (MDR) was found in 9 (42.9%) E. coli and 18 (85.7%) K. pneumoniae isolates; 60% of E. coli and 72.7% of K. pneumoniae were ESBLs producers. blaTEM (50%) and blaCTX-M (30%) were detected in E. coli while blaSHV (83.3%) and blaTEM (55.6%) were detected in K. pneumoniae. One E. coli and two K. pneumoniae isolates harboured NDM-1 gene.
Conclusion: the ASB from this study´s healthy individuals was characterized by MDR bacteria that harboured ESBLs and NDM-1 genes. Thus, emphasizing the need for regular surveillance of bacterial resistance and proper hand hygiene to contain the spread of MDR pathogens in the community.
Enterobacteriaceae are common commensals of the human gastrointestinal tract rendering few benefits to the host as long as they do not acquire virulent traits from invading pathogens in the environment. However, at different anatomical sites, they can become virulent causing varying serious infections which include urinary tract infections (UTIs), sepsis, meningitis, and soft tissue infections depending on their loads and host´s immune system [1]. Notable Enterobacteriaceae that have been commonly implicated in UTIs among humans of all ages and genders are Escherichia coli and Klebsiella pneumoniae [1,2]. UTIs are one of the most common bacterial infections with high economic burden requiring medical care leading to high antibiotic prescription after a visit to a clinician, and occurring more frequently in women than in men because of their shorter urethra that is of closer proximity to the opening of the intestinal tract [3,4]. The high volume of antibiotic prescriptions witnessed in UTIs´ treatment has led to the increasing emergence of antibiotic resistance in the uropathogens causing increased morbidity, delay hospital discharge and expensive cost of treatment which is a serious public health concern [1,5]. Asymptomatic bacteriuria (ASB), a condition that is characterized by the presence of bacteria in two consecutive clear voided urine specimens with both yielding positive cultures of at least 105 CFU/ml of the same uropathogen in an individual without urinary symptoms, occurs at varying frequencies among humans of varying ages but highly prevalent among pregnant women, diabetics and people with genitourinary abnormalities who are more likely to experience symptomatic UTIs along with complications at their much lowered immune status, and can equally become potential transmitters of these causative pathogens to more vulnerable populations in the society [3,6,7]. The UTIs´ aetiologic agents, among which Escherichia coli and Klebsiella pneumoniae are most prevalent, have been reported to be frequently encountered in individuals with ASB [5].
The increasing rate at which Escherichia coli and Klebsiella pneumoniae develop multiple resistance to most of the common antibiotics has seriously hampered the effective treatment of UTIs in both hospital and community settings [1,8,9]. Strains of these organisms that produce extended-spectrum beta-lactamases (ESBLs) have been reported to be generally resistant to many important classes of antibiotics like aminoglycosides and fluoroquinolones besides the beta-lactam antibiotics because of their rapid acquisition of multiple antimicrobial resistance genes on mobile genetic elements aiding their rapid transmission among different bacterial species thereby making UTIs often difficult and expensive to treat [1,8]. The genes encoding the ESBLs production in these bacteria have been identified to be mainly blaTEM, blaSHV and blaCTX-M with varying proportions across different geographical areas of the world [10-12]. The last resort for most ESBLs infections that defer treatment by common antibiotics has been the carbapenems, which are a class of potent beta-lactams with much resistance to most beta-lactamases including ESBLs [13,14]. However, the increasing emergence and rapid dissemination of genes encoding the production of carbapenem-hydrolyzing beta-lactamases among E. coli and K. pneumoniae implicated in UTIs has become one of the most worrisome global public health problems because of the grave consequences of treatment failure such as increased rates of morbidity and mortality [2,9,14,15]. Hence, frequent screening of asymptomatic individuals for surveillance of multiple antimicrobial-resistant uropathogens is therefore, pivotal to current appropriate therapy and strategies for containment of antibiotic resistance in our environments. There is a paucity of data on the study of ASB and the molecular characteristics of its etiologic agents in healthy individuals, even though it is a common occurrence in most African communities. This study, therefore, investigated the phenotypic and molecular characteristics of antibiotic-resistant Escherichia coli and Klebsiella pneumoniae from students with asymptomatic bacteriuria in a Nigerian University, as a means of enriching the antimicrobial resistance surveillance information on ASB from healthy population in this region and thus provides current strategies for UTIs treatment in our environments.
Study design: a cross-sectional study, which aimed at determining the prevalence of ASB and the antibiotic resistance characteristics of common uropathogens among healthy population was carried out among 303 undergraduate students of Niger Delta University, Bayelsa State, as a representative of sexually active individuals in the community. The sample size of this study was determined using the average prevalence of ASB among similar subjects in the previous studies, as described by Charan and Biswas [16].
Study population: the students of various Faculties of Niger Delta University who lived in the hostels within the University environments that had no symptoms or history of UTIs three months before the commencement of the study were randomly drafted into this study from August to November 2019. Niger Delta University is a fast-growing State University situated at Amassoma area of the Wilberforce Island in Southern Ijaw Local Government Area of Bayelsa State on latitude 4�58'13"N longitude 6�06'32" E and 11 meters elevation above the sea level. The students of this University are predominantly from the Southern part of Nigeria.
Inclusion criteria: the students who were not on antibiotics and had no recent history of UTIs were randomly contacted in their various hostels to give their early morning urine samples for research purposes. The interested students gave written informed consent and filled questionnaire to supply their basic demographic data like their Faculty, gender, and age with the exclusion of their names, registration numbers and addresses.
Ethical approval: the permission to conduct this study was obtained from the Ethics and Research Committee of Niger Delta University, Wilberforce Island, Nigeria, before the commencement of the sample collection.
Collection of samples and bacteriology: each volunteer collected a clean mid-stream early morning urine sample into a wide-open sterile universal bottle after being properly instructed on the procedure of collection without touching the skin. The urine samples were labeled appropriately and transported immediately to the laboratory for processing. A 10 �L dispensing loop was used to aseptically transfer each urine sample onto a sterilized nutrient agar plate and properly streaked for discrete colonies before being incubated at 37oC for 24 hours. The sample that had a total of 105 CFU/ml or more was regarded as having significant bacteriuria and the resulting discrete colonies from the plates were separately streaked on HiCromeTM Mueller Hinton Agar, Cysteine lactose electrolyte deficient agar (CLED) and Eosin methylene blue agar (EMB). The isolates with characteristic features on these agar plates were presumptively identified as either E. coli or K. pneumoniae, before storing in glycerol broth at 4oC for further investigations.
Preparation of DNA template for PCR Amplification: the DNA of the isolates was extracted by suspending many colonies from the overnight culture of each isolate on Nutrient agar plates into 100 �L 1X Tris-EDTA buffer, vortexed and boiled at 100oC for 10 minutes as described by Onanuga et al. [17]. The boilate was transferred immediately to the freezer (-20oC) for 10 minutes, maintained at room temperature, vortexed, and centrifuged at 10, 000 rpm for 10 minutes. The resulting supernatant containing DNA of each isolate was collected, stored at 4oC and used as a DNA template for PCR analyses.
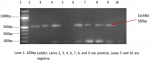
Molecular Identification of E. coli through PCR amplification of 16S rRNA gene: the isolates that gave characteristic colours on EMB, CLED, and Chrom agars were subjected to PCR analysis for the presence of E. coli 16S rRNA (Eco-1) gene as a confirmation for E. coli. The E. coli 16SrRNA gene was screened for in each of the isolate´s DNA by amplifying the gene as previously described by Mamun et al. [18] using the primers in Table 1. The PCR was carried out in a thermal cycler using a 25�l reaction mixture containing 12.5 �l 2x master mix, 0.5 �l each of the forward and reverse primers, 8.5 �l of nuclease-free water and 3 �l of the DNA template at specified conditions in Table 1. The amplified PCR products (10 �l) were evaluated on a 1.5% (w/v) agarose gel at 100 mV for 60 min using BIO-RAD Power Pac 3000 with a molecular weight marker (100 bp DNA Ladder). The DNA bands were then visualized and photographed under UV light after staining the gel with ethidium bromide.
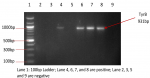
Molecular Identification of K. pneumoniae through PCR amplification of TyrB gene: the isolates that gave characteristic colours on CLED and Chrom agars were subjected to PCR analysis for the presence of the TyrB gene as a confirmation for K. pneumoniae isolates. The TyrB gene was screened for in each of the isolate´s DNA as previously described by Jeong et al. [19] using the primers in Table 1. The K. pneumoniae specific TyrB gene was amplified in the PCR using the same reaction mixture at specified conditions in Table 1. The amplified PCR products were separated on a 1.5% (w/v) agarose gel in the same pattern as described above.
Screening of E. coli and K. pneumoniae isolates for Haemolysin production: haemolytic property of the isolates was determined by inoculating the isolates on freshly prepared sterile 5%v/v Blood agar (consisting of 5 ml of human blood in 100 ml of Nutrient Agar) plates using straight wire loop and incubated at 37oC for 24 hours. Thereafter, the plates were observed for green to black colouration of the agar (partial lysis of red blood cells - alpha haemolysis) and clear zones (complete lysis of red blood cells - beta haemolysis) around inoculated organisms indicating the production of haemolysin.
Screening of E. coli and K. pneumoniae isolates for Biofilm production: biofilm production by the isolates was performed using Congo Red Agar medium which was prepared using the combination of brain heart infusion agar 52 g/L, sucrose 50 g/L and Congo red indicator 8 g/L as described by Mathur et al. [20]. Congo red was prepared as concentrated aqueous solution separately from other medium constituents and then sterilized in different containers before adding them together when the agar had cooled to 55oC before distributing to sterile plates to solidify. The plates were then inoculated with the test organisms and incubated at 37oC for 24 hours before examining for black colonies with a dry crystalline consistency indicating biofilm production.
Antimicrobial Susceptibility Testing and Phenotypic detection of ESBLs: antimicrobial susceptibility testing of the isolates of E. coli and K. pneumoniae was performed using the modified Kirby-Bauer disc diffusion method on Mueller-Hinton agar using commercial discs (Oxoid, UK) against the following antimicrobials; nitrofurantoin (F, 300 μg), cefotaxime (CTX, 30 μg), imipenem (IMP, 10 μg), ceftazidime (CAZ, 30 μg), co-trimoxazole (SXT, 25 μg), gentamicin (CN, 10 μg), chloramphenicol (C, 30 μg) and levofloxacin (LEV, 5 μg) following the Clinical and Laboratory Standards Institute guidelines [21]. The isolates that were resistant to at least one agent in three or more classes of antimicrobial agents used were defined as having multi-drug resistance (MDR) [22]. Furthermore, the isolates that were resistant to Ceftazidime and Cefotaxime were screened for phenotypic production of ESBLs enzymes using combination discs method with discs containing cefpodoxime and cefpodoxime plus clavulanic acid in accordance to CLSI guidelines [21].
Screening for ESBLs genes in the isolates of E. coli and K. pneumoniae: the confirmed isolates of E. coli and K. pneumoniae were screened for the presence of blaTEM, blaSHV and blaCTX-M ESBLs genes as previously described by Bebe et al. [23] and Gholipour et al. [24] using the primers in Table 1. The multiplex PCR reactions were carried out using the same reaction mixture at the specified conditions in Table 1. The amplified PCR products were separated on a 1.5% (w/v) agarose gel in the same pattern as described above.
Screening for Carbapenemases (NDM, VIM, KPC, OXA-48) genes in the isolates of E. coli and K. pneumoniae: the isolates of E. coli and K. pneumoniae that exhibited resistance to Imipenem were screened for the presence of any carbapenemases genes by amplifying NDM, VIM, KPC, OXA-48 genes as described by Balkan et al. [25] and Gokmen et al. [26] using the primers in Table 1. The multiplex PCR reactions were carried out using the same reaction mixture at the specified conditions in Table 1. The amplified PCR products were separated on a 1.5% (w/v) agarose gel in the same pattern as described above.
Statistical analysis: the groups´ differences were tested using the Chi square test (or Fisher´s exact test when expected frequencies were too low), with the assumed level of statistical significance at a P < 0.05 while the strength and direction association or relationship was measured using the phi coefficient/Cramer´s V tests where applicable. Data was performed with SPSS for Windows, version 15.0 (SPSS Inc., Chicago, IL).
A total of 303 urine samples were collected from the healthy University students comprising 144 (47.5%) males and 159 (52.5%) females of ages 15 to 34 years with an average age of 22.7 years. Of all the 303 screened samples, 64 (21.1%) yielded positive significant bacteriuria with a significantly higher prevalence among the females (χ2 = 16.51; P < 0.0001; Odds ratio = 3.459). However, the observed differences in the prevalence of ASB across the age groups are not statistically significant (P > 0.05) as shown in Table 2. Twenty-one (6.9%) E. coli and 21 (6.9%) K. pneumoniae isolates were recovered from the volunteers´ samples, with their respective specific target genes (Figure 1 and Figure 2). The prevalence of E. coli isolates in all the samples was significantly higher in the females (11.3%) than the males (2.1%) (χ2 = 10.0; P = 0.002; Odds ratio = 6). However, the difference observed in the prevalence of K. pneumoniae among the genders was not statistically significant (P = 0.993) as shown in Table 3.
Seven (16.7%) haemolysin producers comprising 6 (28.6%) E. coli and 1 (4.8%) K. pneumoniae were recovered from all the bacterial isolates. Also, 38 (90.5%; 38/42) biofilm producers comprising 17 (81%; 17/21) E. coli and 21 (100%; 21/21) K. pneumoniae were equally recovered from all the bacterial isolates. A total of 4 E. coli and 1 K. pneumoniae isolates produced both haemolysin and biofilm. The antimicrobial susceptibility testing revealed that E. coli and K. pneumoniae isolates generally exhibited higher resistance (57 - 95%) to cefotaxime, ceftazidime, cotrimoxazole, and lower resistance (10 - 33%) to chloramphenicol, levofloxacin, and imipenem. The K. pneumoniae isolates showed higher rates of resistance to most of the tested agents than the E. coli isolates and they were found to exhibit 100% resistance to nitrofurantoin and 71.4% to gentamicin as shown in Table 4. The prevalence of multiple antibiotic resistance (MDR) among the E. coli and K. pneumoniae isolates was 9 (42.9%; 10/21) and 18 (85.7%; 18/21) respectively. The observed difference in the MDR of these organisms is statistically significant (χ2 = 8.4; P = 0.004; Odds ratio = 8). The prevalence of phenotypic ESBLs among the screened isolates of E. coli and K. pneumoniae was 60% and 72.7% respectively. They were all MDR and biofilm-producing isolates.
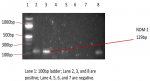
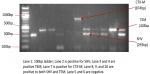
In all, 28 isolates comprising 10 E. coli and 18 K. pneumoniae were screened for the ESBLs genes. A total of 15 (53.6%; 15/28) blaTEM, 15 (53.6%; 15/28) blaSHV and 3 (10.7%; 3/28) blaCTX-M genes were recovered from both organisms, in which E. coli isolates harboured 5 (50%; 5/10) blaTEM and 3 (30%; 3/10) blaCTX-M genes but no blaSHV, whilst K. pneumoniae isolates on the other hand, harboured 10 (55.6%; 10/18) blaTEM and 15 (83.3%; 15/18) blaSHV genes in which 9 (50%; 9/18) of them harboured both blaTEM and blaSHV genes but no blaCTX-M was detected among them (Figure 3). All the organisms´ isolates with the ESBLs genes (24) were resistant to the tested cephalosporins and 22 (91.7%; 22/24) of them were MDR. All the E. coli isolates that expressed phenotypic ESBLs, harboured blaCTX-M genes while 7 (87.5%) of the K. pneumoniae isolates that expressed phenotypic ESBLs, harboured both blaTEM and blaSHV genes while the remaining 1 (12.5%) haboured only blaSHV gene. A total of 3 (60%; 3/5) of the organisms´ isolates that were resistant to imipenem harboured NDM-1 gene (Figure 4). One (50%; ½) was recovered from E. coli isolates while 2 (66.7%; 2/3) were recovered from K. pneumoniae isolates and all exhibited higher levels of MDR harbouring ESBLs genes.
This study´s prevalence of significant bacteriuria among apparently healthy University students was 21.1% with a significantly higher proportion among the females (P < 0.0001) and with no definite association with the age groups of the subjects (P > 0.05). Our prevalence of ASB is similar to the previous findings of Onifade et al. [27] in Ile-Ife, Ngwai et al. [28] in Keffi, Olowe et al. [29] in Oshogbo and Odetoyin et al. [30] in Ile-Ife who reported 20-28.7% rate of ASB among similar Nigerian populations of no overt clinical symptoms. However, the studies of Varli et al. [31] in Turkey and Nsofor et al. [32] in Owerri, Nigeria reported 6.2% and 13.8% lower rate of ASB respectively. This observed difference might be attributed to the sampling techniques, nature, and environments of the populations. Our finding of higher proportion (30.2%) of ASB among the females with a 3.5 times frequent occurrence (greater risk) than the males (P < 0.0001; Odds ratio = 3.5) has been widely reported by previous researchers [23,27-31]. This might be due to the female´s shorter urethra and its closer proximity to the opening of intestinal tract [3,4]. Our result of no definite association of ASB with the age group of the subjects is in agreement with the report of Varli et al. [31] but is in contrast to the findings of Olowe et al. [29] and Odetoyin et al. [30] who reported an increased ASB with increasing age in males. The higher prevalence (30.2%) of ASB among females in this study therefore, indicates the female gender as a major risk factor among healthy populations and this underscores the need for their frequent screening and possible treatment, considering their important roles in the society.
Escherichia coli and K. pneumoniae which were among the leading etiologic agents of UTIs were detected in the same proportion, with E. coli significantly isolated from the females (P = 0.002). The most implicated uropathogen in ASB from healthy populations has been reported by many studies to be E. coli [27-29,31,32] and its higher prevalence among females has equally been reported by Ngwai et al. [28]. However, the recovery of K. pneumoniae in equal proportion with E. coli in this study suggests the increasing rate at which K. pneumoniae is becoming an important agent of ASB which might result into symptomatic UTIs in humans. This is in agreement with the findings of Onanuga and Selekere, [33] in Bayelsa State who reported K. pneumoniae as the most implicated uropathogen in a similar ASB study. This therefore, emphasizes the need for microbial culture identification for better management of UTIs since different organisms behave differently to treatments.
Haemolysin production by certain strains of bacteria has been attributed to the secretion of alpha-haemolytic toxin which is associated with significant disease burden in healthy humans [34,35]. The screening for haemolysin production among this study´s bacteria revealed a total prevalence of 7 (16.7%) comprising 6 (28.6%) E. coli and 1 (4.8%) K. pneumoniae isolates, with a higher proportion of 6 (85.7%) recovered from the females. The observed low level of these toxin-producing isolates suggests that the bacteria emanated from healthy individuals without overt symptoms. However, it also suggests that the females in this study might be highly predisposed to E. coli associated UTIs in the event of a lowered immune status and might constitute a transmission agent to the vulnerable healthy community. A high prevalence of 38 (90.5%) biofilm producers comprising 17 (81%) E. coli and 21 (100%) K. pneumoniae isolates were recovered in this study. This therefore, suggests the possibility of a high-level multidrug resistance tendency among the isolates since biofilm-producing bacteria have been reported to exhibit varying degrees of resistance to antimicrobial agents and host defense strategies, thereby causing a serious public health problem of treatment failures [35-37]. This observation is an indication that these organisms might have emanated from the population areas where antimicrobials have been grossly misused which therefore support the postulation that healthy members of the population are the highest reservoirs of antimicrobial-resistant bacteria [38,39].
This study´s antimicrobial susceptibility test results revealed that the organisms´ isolates generally exhibited high in vitro resistance of 57 - 95% to cefotaxime, ceftazidime, and co-trimoxazole, a finding that has been widely reported in Gram-negative bacilli from ASB [27,33]. This may be attributed to the extensive use of these agents in the treatment of uncomplicated UTIs in both hospital and community settings since extensive use of antimicrobial agents favours the selection of antibiotic-resistant bacteria and hence, a major risk factor for antibiotic resistance in a community [30,40]. This therefore, implies that these agents should no longer be considered for the empiric therapy of UTIs. The E. coli isolates in this study showed low to moderate (28.6 - 33.3%) in vitro resistance to gentamicin, chloramphenicol, and nitrofurantoin but K. pneumoniae exhibited higher in vitro resistance to gentamicin (71%) and nitrofurantoin (100%). Our result also reveals that K. pneumoniae generally exhibited higher resistance to all the tested agents than E. coli which is in agreement with the reports of previous studies in Africa [2,41]. The most effective agents in this study are levofloxacin and imipenem since the organisms exhibited very low resistance to them. This data therefore, highlights the importance of routine antimicrobial susceptibility testing as a guide to improving infection control and the use of antimicrobial agents in the community.
The prevalence of multiple drug resistance in this study was E. coli (42.9%) and K. pneumoniae (85.7%) which show a significantly higher MDR among K. pneumoniae isolates with 8 times frequent occurrence (greater risk) than E. coli (P = 0.004; χ2 = 8.4; Odds ratio = 8) which implies that these organism´ isolates might have been exposed to many classes of antibiotics hence possessing many antibiotic resistance genes and this has been ascribed to the widespread use of antimicrobial agents in both agricultural and health care settings [2]. The observed higher MDR in K. pneumoniae may be attributed to its higher biofilm-forming ability over E. coli as revealed in this study and which has been reported to be a major risk factor for multidrug resistance [35-37]. This non-susceptibility of urinary K. pneumoniae to most commonly used antibiotics is a global health problem that is rapidly increasing with varying dimensions across regions which call for regular surveillance programs to monitor its resistance and spread [2]. ESBLs producing strains of bacteria that were previously associated with nosocomial UTIs were now detected in 60% of E. coli and 72.7% of K. pneumoniae screened isolates in this study. These isolates were all observed to be MDR and biofilm producers, implying that the ESBL enzymes which were primarily produced to hydrolyze beta-lactams antibiotics are mediated by plasmids which also harbour other classes of antibiotics resistance determinants thereby making the bacteria to be non-susceptible to most commonly used antibiotics [42-44]. The detection of a high proportion of this type of bacteria among healthy individuals in a community setting may pose a serious health problem of therapeutic difficulties raising major concern.
The screening for ESBLs genes among these study´s isolates, which is the first report of its kind in Nigeria, revealed a total prevalence of 15 (53.6%) blaTEM, 15 (53.6%) blaSHV and 3 (10.7%) blaCTX-M genes among the screened E. coli and K. pneumoniae isolates that largely exhibited MDR and expressed phenotypic ESBLs. This finding is similar to the reports of Ojdana et al. [45] and Hamad and Mustafa, [46] from hospital-based urine samples in Poland and Iraq respectively. These beta-lactamases genes that encode antibiotic resistance to broad-spectrum beta-lactam drugs have been reported to be located on transferable genetic elements such as plasmids or transposons which can facilitate the horizontal transmission of resistance among other bacteria species [44,45,47].This therefore suggests a possibility of rapid spreading of multi-resistance among bacteria within the host´s community since the isolates harbouring them were MDR.
The gene encoding the New Delhi Metallo-beta-lactamase-1 (NDM-1), a carbapenemase beta-lactamase enzyme of the class B Metallo-beta-lactamase, was screened for in 2 E. coli and 3 K. pneumoniae isolates that exhibited in vitro resistance to imipenem in this study. A total of 3 (7.1%; 3/42) isolates comprising E. coli (1) and K. pneumoniae (2), which exhibited a high level of MDR and possessed ESBLs genes, harboured the NDM-1 gene. The bacteria strains that showed resistance to carbapenems, one of the last resort agents that are used in treating multidrug-resistant ESBLs producing bacterial infections have been reported to be difficult to treat, causing increased death in hospital patients that are infected by them [48,49]. This NDM-1 gene whose emergence has been ascribed to the excessive use of antibiotics, is located on the bacterial plasmids and can be spread from one strain of bacteria to another by horizontal transfer thus, accounts for its increasing emergence across the globe even beyond the hospital settings [49,50]. Hence, the detection of this gene among isolates from healthy individuals is therefore a serious threat to public health which calls for concerted efforts by all health stakeholders to contain its spread in the community through education on the rational use of antibiotics and strict personal proper hand hygiene.
The study reports a higher burden of ASB among healthy University students that is characterized with high proportions of MDR E. coli and K. pneumoniae isolates that harboured ESBLs and NDM-1 genes thereby predicting a likely outbreak of community-associated UTIs that will be difficult to treat. This therefore, calls for regular surveillance of bacterial resistance coupled with education on prudent use of antibiotics and strategies to contain the spread of MDR pathogens in our community.
What is known about this topic
- Asymptomatic Bacteriuria (ASB) is commonly encountered among healthy females´ subjects than the males;
- Escherichia coli strains are the most prevalent uropathogens that are associated with ASB in healthy individuals which may result into urinary tract infections;
- Treatment of UTIs arising from ASB can be achieved with most commonly available antibiotics in the community with a lower incidence of MDR among the uropathogens.
What this study adds
- Escherichia coli and Klebsiella pneumoniae isolates were recovered in equal proportion in the ASB from healthy subjects, and the proportion of haemolysin-producing bacteria was higher among the Females subjects than the males;
- The ESBLs genes mainly blaTEM, blaSHV and blaCTX-M genes were detected among the bacterial isolates with higher levels of MDR;
- The detection of blaNDM-1 gene, a carbapenemases plasmid encoding gene in the urinary E. coli and K. pneumoniae strains from healthy individuals, is the first report in Nigeria.
The authors declare no competing interests.
Adebola Onanuga participated in designing the study, performed the experiments, analyzed the data and wrote the manuscript. Darlington Deboh Eboh participated in the experiments and review of the manuscript for publication. Babatunde Odetoyin performed the molecular experiments, participated in data analysis and review of the manuscript for publication. Ocholi Jonathan Adamu participated in the review of the manuscript for publication. All authors read and approved the final manuscript.
The authors thank the technologists of the laboratories for their support and assistance during the period of the study.
Table 1: primers used for PCR amplification and sequencing
Table 2: the distribution of significant bacteriuria among the subjects´ age groups
Table 3: the prevalence of E. coli and K. pneumoniae among the healthy volunteers
Table 4: antimicrobial resistance patterns of the volunteers´ urinary bacteria isolates
Figure 1: amplification of 16sRNA of E. coli
Figure 2: amplification of TyrB of Klebsiella pneumoniae
Figure 3: amplification of SHV, TEM, CTX-M
Figure 4: amplification of NDM 1
- Tansarli G, Athanasiou S, Falagas M. Evaluation of Antimicrobial Susceptibility of Enterobacteriaceae Causing Urinary Tract Infections in Africa. Antimicrobial Agents and Chemotherapy. 2013 Aug;57(8):3628-39. PubMed | Google Scholar
- El Bouamri MC, Arsalane L, El Kamouni Y, Zouhair S. Antimicrobial susceptibility of urinary Klebsiella pneumoniae and the emergence of carbapenem-resistant strains: a retrospective study from a university hospital in Morocco, North Africa. African Journal of Urology. 2015;2(1):36-40. Google Scholar
- Demilie T, Beyene G, Melaku S, Tsegaye W. Urinary bacterial profile and antibiotic susceptibility pattern among pregnant women in North West Ethiopia. Ethiop J Health Sci. 2012 Jul;22(2):121-8. PubMed | Google Scholar
- Abbo LM, Hooton TM. Antimicrobial stewardship and urinary tract infections. Antibiotics. 2014 May 5;3(2):174-92. PubMed | Google Scholar
- Melaku S, Kibret M, Abera B, Gebre-Sellassie S. Antibiogram of nosocomial urinary tract infections in Felege Hiwot referral hospital, Ethiopia. Afr Health Sci. 2012; 12(2):134-9. PubMed | Google Scholar
- Colgan R, Nicolle LE, McGlone A, Hooton TM. Asymptomatic bacteriuria in adults. Am Fam Physician. 2006 Sep 15;74(6):985-90. PubMed | Google Scholar
- Girishbuabu RJ, Srikrishna R, Ramesh ST. Asymptomatic bacteriuria in pregnancy. Inter. J Biology & Medical Research. 2011;2(3):740-742. Google Scholar
- Canton R, Ruiz-Garbajosa P. Co-resistance: an opportunity for the bacteria and resistance genes. Curr Opin Pharmacol. 2011 Oct;11(5):477-85. PubMed | Google Scholar
- Eshetie S, Unakal C, Gelaw A, Ayelign B, Endris M, Moges F. Multidrug resistant and carbapenemase producing Enterobacteriaceae among patients with urinary tract infection at referral Hospital, Northwest Ethiopia. Antimicrobial Resistance and Infection Control. 2015; 4:12. PubMed | Google Scholar
- Muhammad MH, Swedan S. Molecular and Phenotypic Characterization of Carbapenem Resistance and Extended Spectrum Beta-Lactamases among Urinary Escherichia coli Isolates. Inter J Sci and Technol. 2015; 5(9). PubMed | Google Scholar
- Rani S, Jahnavi I, Nagamani K. Phenotypic and Molecular Characterization of ESBLs producing Enterobacteriaceae in A Tertiary Care Hospital. IOSR Journal of Dental and Medical Sciences. 2016; 15(8):27-34. Google Scholar
- Zorgani A, Almagatef A, Sufya N, Bashein A, Tubbal A. Detection of CTX-M-15 Among Uropathogenic Escherichia coli Isolated from Five Major Hospitals in Tripoli, Libya. Oman Medical Journal. 2017; 32(4):322-327. PubMed | Google Scholar
- Breilh D, Texier-Maugein J, Allaouchiche B, Saux MC, Boselli E. Carbapenems. J Chemother. 2013; 25(1):1-17. PubMed | Google Scholar
- Huttner A, Harbarth S, Carlet J, Cosgrove S, Goossens H, Holmes A et al. Antimicrobial resistance: a global view from the 2013 World Healthcare-Associated Infections Forum. Antimicrob Resist Infect Control. 2013; 2:31. PubMed | Google Scholar
- Nordmann P. Carbapenemase-producing Enterobacteriaceae: overview of a major public health challenge. Med Mal Infect. 2014 Feb;44(2):51-6. PubMed | Google Scholar
- Charan J, Biswas J. How to calculate sample size for different study designs in medical research. Indian J Psychological Medicine. 2013;35(2):121-126. PubMed | Google Scholar
- Onanuga A, Igbeneghu O, Lamikanra A. A Study of the Prevalence of Diarrhoeagenic Escherichia coli in Children from Gwagwalada, Federal Capital Territory, Nigeria. Pan African Medical Journal. 2014;17:146. PubMed | Google Scholar
- Mamun MM, Parvej MS, Ahamed S, Hassan J, Nazir KHMNM, Nishikawa Y et al. Prevalence and Characterization of Shiga Toxigenic Escherichia coli in Broiler Birds in Mymensingh. Bangl J Vet Med. 2016; 14(1):5-8. Google Scholar
- Jeong E, Lee K, Heo S, Seo J, Choi Y. Rapid Identification of Klebsiella pneumoniae, Corynebacterium kutscheri, and Streptococcus pneumoniae Using Triplex Polymerase Chain Reaction in Rodents. Exp Anim. 2013; 62(1):35-40. PubMed | Google Scholar
- Mathur T, Singhal S, Khan S, Upadhyay DJ, Fatma T, Rattan A. Detection of Biofilm formation among the Clinical isolates of Staphylococci: An evaluation of three different screening methods. Indian Journal of Medical Microbiology. 2006;24(1):25-9. PubMed | Google Scholar
- Clinical and Laboratory Standards Institute (CLSI). Performance Standards for antimicrobial susceptibility testing. Twenty seventh Information Supplement. 2017; M100-S27:62-71.
- Magiorakos AP, Srinivasan A, Carey RB, Carmeli Y, Falagas ME, Giske CG et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect. 2012; 18(3):268-281. PubMed | Google Scholar
- Bebe T, Odetoyin B, Bolarinwa R. Occurrence of Multidrug-resistant Uropathogens Implicated in Asymptomatic Bacteriuria in Adults with Sickle Cell Disease in Ile-Ife, Southwest Nigeria. Oman Med J. 2020;35(2):e109. PubMed | Google Scholar
- Gholipour A, Soleimani N, Shokri D, Mobasherizadeh S, Kardi M, Baradaran A. Phenotypic and Molecular Characterization of Extended-Spectrum β-Lactamase Produced by Escherichia coli, and Klebsiella pneumoniae Isolates in an Educational Hospital. Jundishapur Journal of Microbiology. 2014; 7(10):e11758. PubMed
- Balkan II, Aygun G , Aydin S, Mutcali SI, Kara Z, Kuskucu M et al. Blood stream infections due to OXA-48-like carbapenemase-producing Enterobacteriaceae: treatment and survival. Inter J Infectious Dis. 2014; 26:51-56. PubMed | Google Scholar
- Gokmen TG, Nagiyev T, Meral M, Onlen C, Heydari F, Koksal F. NDM-1 and rmtC-Producing Klebsiella pneumoniae isolates in Turkey, Jundishapur. J Microbiol. 2016; 9(10):e33990. PubMed | Google Scholar
- Onifade AK, Anibijuwon II, Azariah EJ. Urinary tract infection in apparently healthy individuals in Ile-Ife, Nigeria: detection of predominant microorganisms and antibiotics susceptibility profile. Afri J Microbiol Res. 2011; 5(20):3233-36. Google Scholar
- Ngwai YB, Iliyasu H, Young E, Owuna G. Bacteriuria and antimicrobial susceptibility of Escherichia coli isolated from urine of Asymptomatic University Students in Keffi, Nigeria. Jundishapur J Microbiol. 2012;5(1):323-7. Google Scholar
- Olowe OA, Makanjuola OB, Olabiyi KO, Akinwusi PO, Alebiosu CO, Isawumi MA et al. Asymptomatic bacteriuria among elderly and middle aged rural community dwellers in South Western Nigeria. Infect Drug Resist. 2013;6:55-8. PubMed | Google Scholar
- Odetoyin BW, Olaniran O, Afolayan D, Aderibigbe I, Alaka O, Onanuga AA. Asymptomatic Bacteriuria In An Apparently Healthy Population And Its Relation To Hypertension. Afri J Clin. Experimental Microbiol. 2018; 19(4):282-290. Google Scholar
- Varli M, Guruzb H, Arasa S, Yalcina A, Atlia T, Turgaya M. Asymptomatic bacteriuria among the elderly living in the community: prevalence, risk factors and characteristics. Eur Geriatr Med. 2012;3(2):87-91. Google Scholar
- Nsofor CA, Obijuru CE, Ozokwor CL. Asymptomatic Bacteriuria Among Female Students of a Tertiary Institution in Southeast Nigeria. EC Bacteriology and Virology Research. 2016;2(3):106-112.
- Onanuga A, Selekere TL. Virulence and antimicrobial resistance of common urinary bacteria from asymptomatic students of Niger Delta University, Amassoma, Bayelsa State, Nigeria. J Pharmacy Biomedical Sci. 2016;8(1):29-33. PubMed | Google Scholar
- Berube BJ, Wardenburg JB. Staphylococcus aureus α-Toxin: Nearly a Century of Intrigue. Toxins (Basel). 2013; 5(6):1140-1166. PubMed | Google Scholar
- Karam MRA, Habibi M, Bouzari S. Relationships between Virulence Factors and Antimicrobial Resistance among Escherichia coli Isolated from Urinary Tract Infections and Commensal Isolates in Tehran, Iran. Osong Public Health Res Perspect. 2018; 9(5):217-224. PubMed | Google Scholar
- Archer NK, Mazaitis MJ, Costerton JW, Leid JG, Powers ME, Shirtliff ME. Staphylococcus aureus biofilms Properties, regulation and roles in human disease. Virulence. 2011; 2(5):445-459. PubMed | Google Scholar
- de la Fuente-Nunez C, Reffuveille F, Fernandez L, Hancock RE. Bacterial biofilm development as a multicellular adaptation: antibiotic resistance and new therapeutic strategies. Curr. Opin. Microbiol. 2013 Oct;16(5):580-9. PubMed | Google Scholar
- Lamikanra A, Ako-Nai Ak, Ogunniyi DA. Transferable antibiotic resistance in Escherichia coli isolated from healthy Nigerian school children. Int J Antimicrob Agents. 1996 May;7(1):59-64. PubMed | Google Scholar
- Onanuga A, Oyi AR, Olayinka BO, Onaolapo JA. Prevalence of community-associated multi-resistant Staphylococcus aureus among healthy women in Abuja, Nigeria. African Journal of Biotechnology. 2005; 4(9):942-945. Google Scholar
- Abejew AA, Denboba AA, Mekonnen AG. Prevalence and antibiotic resistance pattern of urinary tract bacterial infections in Dessie area, North-East Ethiopia. BMC Research Notes. 2014;7:687. PubMed | Google Scholar
- Shatalov A. Prevalence and Antibiotic Resistance Pattern of Escherichia coli and Klebsiella pneumoniae in Urine Tract Infections at the La Paz Medical Center, Malabo, Equatorial Guinea. Open J. Medical Microbiol. 2015; 4(5):177-183. Google Scholar
- Canton R, Coque TM. The CTX-M β-lactamase pandemic. Curr Opin Microbiol. 2006 Oct;9(5):466-75. PubMed | Google Scholar
- Charfi K, Grami R, Ben-Jeddou A, Messaoudi A, Mani Y, Bouallegue O et al. Extended-spectrum β-lactamases and plasmid-mediated quinolone resistance in enterobacterial clinical isolates from neonates in Tunisia. Microb Pathog. 2017;110:184-188. PubMed | Google Scholar
- Adamus Bialek W, Baraniak A, Wawszczak M, Gluszek S, Gad B, Wróbel K et al. The genetic background of antibiotic resistance among clinical uropathogenic Escherichia coli strains. Molecular Biology Reports. 2018 Oct;45(5):1055-1065. PubMed | Google Scholar
- Ojdana D, Sacha P, Wieczorek P, Czaban S, Michalska A, Jaworowska J et al. The occurrence of blaCTX-M, blaSHV, and blaTEM genes in extendedspectrum β-lactamase-positive strains of Klebsiella pneumoniae, Escherichia coli, and Proteus mirabilis in Poland. Int J Antibiot. International Journal of Antibiotics. 2014;2014:935842. Google Scholar
- Hamad PA, Mustafa KK. Prevalence of blatem, blashv, and blactx-M genes among ESBL-Producing Klebsiella pneumoniae and Escherichia coli isolated from Thalassemia in Erbil, Iraq. Mediterr J Hematol Infect Dis. 2019; 11(1): e2019041. PubMed | Google Scholar
- El salabi A, Walsh TR, Choichanic C. “Extended spectrum β-lactamases, carbapenemases and mobile genetic elements responsible for antibiotics resistance in Gram-negative bacteria.” Critical Reviews in Microbiology. 2012; 39(2):113-122. Google Scholar
- Kumarasamy KK, Toleman MA, Walsh TR, Bagaria J, Butt F, Balakrishnan R et al. Emergence of a new antibiotic resistance mechanism in India, Pakistan, and the UK: a molecular, biological, and epidemiological study. Lancet Infect Dis. 2010;10(9):597-602. PubMed | Google Scholar
- Khan AU, Maryam L, Zarrilli R. Structure, Genetics and Worldwide Spread of New Delhi Metallo-β-lactamase (NDM): a threat to public health. BMC Microbiol. 2017 Apr 27;17(1):101. PubMed | Google Scholar
- Hudson CM, Bent ZW, Meagher RJ, Williams KP. Resistance Determinants and Mobile Genetic Elements of an NDM-1-Encoding Klebsiella pneumoniae Strain. PLOS One. 2014; 9(6):e99209. PubMed | Google Scholar