A narrative review of dengue fever infection and epidemic activity in Kenya (2010 to 2020)
Carren Bosire, Francis Mutuku, Bryson Ndenga, Makenzi Nzaro, Kavinya Mwendwa, Angelle Desiree LaBeaud
Corresponding author: Carren Bosire, Department of Pure and Applied Sciences, Technical University of Mombasa, Mombasa, Kenya 
Received: 14 May 2023 - Accepted: 25 Sep 2023 - Published: 16 Oct 2023
Domain: Community health,Health Research,Public health
Keywords: Dengue virus, seroprevalence, epidemics, Kenya
©Carren Bosire et al. PAMJ-One Health (ISSN: 2707-2800). This is an Open Access article distributed under the terms of the Creative Commons Attribution International 4.0 License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Cite this article: Carren Bosire et al. A narrative review of dengue fever infection and epidemic activity in Kenya (2010 to 2020). PAMJ-One Health. 2023;12:10. [doi: 10.11604/pamj-oh.2023.12.10.40416]
Available online at: https://www.one-health.panafrican-med-journal.com/content/article/12/10/full
A narrative review of dengue fever infection and epidemic activity in Kenya (2010 to 2020)
![]() Carren Bosire1,&, Francis Mutuku2,
Carren Bosire1,&, Francis Mutuku2, ![]() Bryson Ndenga3, Makenzi Nzaro1,
Bryson Ndenga3, Makenzi Nzaro1, ![]() Kavinya Mwendwa1, Angelle Desiree LaBeaud4
Kavinya Mwendwa1, Angelle Desiree LaBeaud4
&Corresponding author
Introduction: dengue cases occur frequently in Kenya, although the trend is not uniform across the country, making this infectious disease a major public health problem that is transmitting without detection. The purpose of this review study was to determine the seroprevalence of dengue virus (DENV) infection in the Kenyan population during the last ten years to reveal geographical areas with unreported dengue fever activity. For that purpose, this review describes the recent (2010 to 2020) DENV incidences, methods of detection, and the circulating genotypes in Kenya.
Methods: studies were identified to summarize reported dengue fever infections and epidemic activities in Kenya using a search approach (terms “Dengue”, “Dengue Virus” and “DENV” in combination with “Infection in Kenya”). All published studies between 2010 and 2020 retrieved through search engines PubMed, Google Scholar, Research4life, and ScienceDirect were screened for eligibility.
Results: a total of 21 eligible articles containing 24 studies, covering five general geographic regions and 39 sampling sites, were included in this review. The results indicate a surveillance gap in dengue fever infection serosurvey in Kenya. There was transmission between epidemics in the reported areas. Seroprevalence ranged from 0.4% to 100% during outbreaks. Coast region was the most studied region with Mombasa being the most sampled site.
Conclusion: about 76.6% of Kenyan Counties' DENV seroprevalence status was found to be unknown or unreported. In addition, DENV infection was unrecognized and/or unreported in most areas of the country, especially rural Kenya. This information may serve as a basis for better awareness and detection of DENV infection during outbreaks and in establishing appropriate prevention and control measures to further avoid outbreaks.
Dengue virus (DENV) is an arthropod-transmitted, positive stranded RNA virus of the family Flaviviridae and the genus flavivirus. It is delineated into four serotypes (DENV-1 through DENV-4), all of which are transmitted by Aedes mosquitos, mainly Aedes aegypti and to a lesser extent by Aedes albopictus [1]. Each of the four serotypes has ability to cause self-limiting fevers and fatal conditions such as dengue hemorrhagic fever (DHF) and dengue shock syndrome (DSS) [2].
Globally, DENV infections are estimated to be 100 to 400 million infections per year. DENV is endemic in more than 100 countries in Africa, America, Eastern Mediterranea, South-East Asia, and Western Pacific, with Asia representing approximately 70% of the global disease burden. It is reported to be spreading to previously unentered regions such as Europe, France, Croatia, and Afghanistan [3]. Reported cases have increased from 505,430 in 2000 to 5.2 million in 2019 [4]. Brady et al. model estimate 390 million DENV infections per year, out of which 96 million manifest clinically [5]. Africa has reported 45 DENV outbreaks between 1960 and 2020, out of which 17 occurred in East Africa and 16 in West Africa. About 2211 cases were reported in the same period, with 1954 (88.4%) reported during outbreaks [2]. Kenya has experienced substantial DENV infections and outbreaks since 1982 [6-10]. Climate change is associated with changes in seasonal weather patterns with subsequent impacts on the suitability and temporal as well as spatial distribution of these infections [11].
DENV infection is a major world public health problem [3], and Kenya has not been spared, reporting an enormous burden. A study among children with undifferentiated febrile illness in Kenya reported more than 40% infection out of 1,022 febrile children [12]. In another study, 40% (345/868) of febrile patients tested positive for dengue [6]. Additionally, during 2017 DENV outbreak, 56% (672/1,199) of suspected patients were DENV positive [13]. Although disease manifestation during the outbreaks have not been reported to be severe, DENV infection complications progressing to dengue hemorrhagic fever and deaths have been reported [9,13,14].
Precipitous climate change, uncontrolled urbanization, travelling within endemic areas, movement of populations, and massive deforestation are among several factors that have been associated with flavivirus infection and transmission [15]. Although reliance on household water storage has been associated with increased exposure to flaviviruses [16], and transmission determined by vector abundance which differs regionally in Kenya [17,18], a better clinical characterization of these infections is desirable to improve the quality of care given to patients [19]. Reported DENV outbreak challenges include limited laboratory capacity for testing suspected samples, limited essential supplies for dengue fever diagnosis, confirmation, and treatment, and weak rapid response capacity. The latter is occasioned by limited training, yet a multidisciplinary team of experts is required for an effective response to an outbreak [13].
The endemicity of DENV in Kenya is undisputable [1,9], but the prevalence data currently available does not accurately represent the severity of exposure, infection, disease, and its variation by region [6,20]. There is more dengue in Kenya than indicated by public health reporting [21], resulting in scanty information available regarding the incidence of dengue across Kenya. DENV infections in Kenya could be unrecognized and misdiagnosed [12,14,22,23]. In addition, dengue virus could be circulating at low levels in most areas in Kenya, with a possibility of an outbreak any time. In a case scenario, studies in unreported non-epidemic settings have reported DENV seropositivity [23]. To fill the surveillance gap, areas in the country where testing has never been carried out need to be revealed. This can inform national control strategies to consider these areas as priority for surveillance to put in place preventive measures to avert future outbreaks. In addition, there is need to detect the burden of DENV disease throughout the country to provide effective control measures targeting appropriate vectors. This can be done by mapping out neglected, unreported and non-epidemic areas which are potential “breeding grounds” for DENV outbreaks.
Therefore, to redress the dearth of Kenyan DENV regional seroprevalence information, the objective of this review was to describe the recent (2010 to 2020) DENV incidences and epidemic activities, methods of DENV detection, and the circulating genotype in humans. The information may serve as a basis for better awareness and detection of DENV infection during outbreaks and in establishing appropriate prevention and control measures. The study narrowed to DENV infection in the last decade (2010 - 2020) since prior to that, there were limited reports on DENV infection in Kenya. The first recorded case was in 1982 reported in Malindi and Kilifi Counties at the Kenyan coast involving DENV-2 [24], and the same serotype was again reported in Kilifi much later in 1997 [25]. But in the last decade, several outbreaks have been reported [6,9,13,14,26-28], probably due to improved diagnosis, more surveillance, and increased spread of the virus because of human movement from endemic to non-endemic areas.
Search strategy and selection criteria
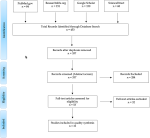
We carried out a search for all published studies on DENV in Kenya conducted between 2010 and 2020, retrieved through search engines PubMed, Google Scholar, Research4life, and ScienceDirect, with reference to the Preferred Reporting Items for Systematic reviews and Meta-Analysis (PRISMA) guidelines where applicable [29], with our inclusion and exclusion criteria (Table 1). The databases were queried to search the titles and abstracts using the keywords “Dengue”, “Dengue Virus” and “DENV” in combination with “Infection in Kenya”. The “publication date” filter was set at 2010 - 2022. Identified relevant studies were screened and filtered for eligibility using a four- step search strategy of PRISMA (Figure 1), i.e., identification, screening, eligibility, and inclusion of studies.
Data analysis
All eligible full articles were analyzed for DENV infection and/or epidemic activity capturing information regarding the broad regional set-up, study site, period of study, study population, sample size, number of DENV positive cases, case identification procedure, serotype and genotype, and study ID.
Quality assessment
The methodological quality of selected DENV infection seroprevalence studies was evaluated by two reviewers using a quality assessment checklist adapted from Hoy et al. [30]. Risk of bias was assessed using nine domains with risk of bias levels being low (score = 0) or high (score = 1), and the overall risk of bias defined as low (score 0-3), moderate (4-6), and high (7-9). Discrepancies between the two scores were resolved through discussion with a third reviewer. Both assessors rated 10 studies as low risk of bias, and 14 studies as moderate risk of bias. Overall risk of bias was low (3) (Table 2).
Four hundred and fifty-three (453) records from PubMed (84), ScienceDirect (68), Research4life (151), and Google Scholar (150) were identified through database search. The latter had an enormous volume of records (11,801) necessitating narrowing the search selection to “relevance” to research articles only without reviews, finally yielding the 150 articles. After de-duplication, a total of 337 original titles were selected for abstract review. The abstract review resulted in the exclusion of 284 reports, remaining with 53 full text articles for assessment for eligibility. Out of these, 32 full text articles were further excluded for several reasons. To highlight the reasons, some articles were pre-prints or poster presentations; others were generalized studies on flavivirus not specific to DENV. Also, some articles were retrospective studies done in the period 2010-2020, but using data collected outside the study period. In addition, there were articles evaluating economic burden of DENV infection, characteristics of DENV vectors, single DENV patient case study, non-human DENV infection, DENV infection in vector mosquitoes, DENV infection manifestations, dengue vector entomological studies among others. Assessment was conducted on a total of 21 remaining eligible articles (Figure 1), which satisfied our selection criteria (Table 1).
Reported DENV infections (2010 - 2020)
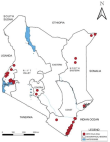
The 21 eligible full text articles for analysis contained 24 studies, covering 5 broad geographical regions in Kenya, with 39 sampling sites. Of the 5 regions, Coast was the most studied with 21 studies, followed by Western (12), North-Eastern (4), Central (2), while Rift Valley was the least studied (1). Most of the sampling sites were studied more than once (Table 3), with both Mombasa and Chulaimbo having the highest (5 and 4 respectively). Of the 47 counties in Kenya, DENV seroprevalence was studied in only 11 Counties (23.4%). These were 5 counties in the Coast region (Mombasa, Kilifi, Lamu, Taita-Taveta, and Kwale), 2 in Western Kenya (Kisumu and Busia), 2 in North-Eastern (Mandera and Wajir), 1 in Rift Valley (Trans-Nzoia), and 1 in Central (Nairobi) (Figure 2). There were 24 study periods, with a lot of overlap, from 2010 to 2019. Most studied periods fell in 2014 (11), while others were 2010 (5), 2011 (7), 2012 (1), 2013 (1), 2015 (4), 2016 (4) 2017 (6), 2018 (1), and 2019 (1). Of the 24 studies analyzed in this review, 18 were carried out during inter-epidemic DENV transmission periods (Table 3).
Reported DENV Epidemics (2010-2020)
There were eight sporadic outbreaks reported during the study period, including 2 reported by WHO [13] (Table 4). Of the 7 that reported causative serotype, DENV-2 featured in 5 outbreaks, the exceptions being DENV-3 outbreaks in North-Eastern in 2011 [9] and South Coast (Mtongwe) in March 2019 [27]. DENV-3 and DENV-1 featured in 4 and 2 outbreaks, respectively. Disease manifestation during the outbreaks were not severe although there were reports of hemorrhagic manifestations [9,14], and two fatal cases [13,14]. Co-infection with more than one serotype was detected is some outbreaks involving DENV-2 and 3 [3], and DENV-1 and 2 [12]. Twenty three percent (23%) mixed infection with malaria among laboratory-confirmed dengue patients was also reported in another study [9]. Reported interventions included supply of rapid dengue diagnostic kits to local health clinics and dispatch of response teams tasked with initiating community sensitization on infection prevention, mosquito control activities [6], and indoor as well as outdoor residual spraying by health authorities [9].
Methods of DENV detection
In this study, there were three study designs: community-based serosurveys, health facility-based surveillance, and retrospective studies. Of the 24 study periods, 17 designs were health facility-based fever serosurveys, 4 community-based serological surveys, and 3 retrospective studies. The total sample size studied was 20,885 individuals with 3,092 DENV positive cases. Data for ratio of male to female positive cases was incomplete, but from the studies that were complete, health facility-based studies had higher females than males, whereas serological surveys had more males than females. Generally, DENV infectivity in children increased with age. In cross-sectional studies, young adults showed higher DENV infectivity. Weighted seroprevalence was calculated as percentage of positive DENV samples out of sample size. It ranged from 0.45% in a health facility-based fever surveillance [16], to 100% in two outbreak studies where all the 14 and 10 sampled patients tested positive for DENV [10,26] (Table 3).
There was a wide range of DENV identification procedures, with two most used procedures being ELISA-IgG and Reverse transcription PCR (RT-PCR), used 14 and 12 times respectively. The latter included real time reverse transcription PCR (RT-PCR) and Quantitative real time reverse-transcription PCR (qRT-PCR). A particular method was used as the only identification procedure and/or in combination with other identification procedures. Most of the studies in this paper (12 out of 20) used a combination of anti-DENV IgM/IgG/IgA ELISAs and NS1 ELISA, or a combination of diagnostic tests (Table 5). Notably, study reports in this paper emphasized the need for a constant review and making available comprehensive differential disease diagnostic panels at health facilities and testing laboratories. This is feted to improve case detection and clinical diagnosis.
Circulating DENV genotypes
All DENV 1-4 serotypes and their representative lineages were identified by different studies in this analysis (Table 6). In the entire study, there were 42 DENV serotype occurrences, with DENV-2 being the most dominant (18), followed by DENV-3 (10), DENV-1 (9), and the least was DENV-4 (5). DENV-1 lineage was related to Djibouti 1998 strain genotype I, and Thailand 1964 strain genotype II (related to Indonesia 2012 strain genotype II). DENV-2 was African genotype IV strain, C-I lineage (India 2011), C-II lineage (Burkina Faso 1979 - 2013), and cosmopolitan genotype (India 2009-2010, 2014). DENV-3 was genotype III (Pakistan 2007-2009), genotype V (related to Brazil 2006), genotype V strain (Philippines 1956), and genotype III strain (related to Pakistan 2006, China 2013, and India 2009). DENV-4 was genotype II strain (related to Indonesia 2012) (Table 6).
The present study evaluated seroprevalence of DENV infection in Kenya, an example of flavivirus infection that represent a major global health problem. All DENV 1-4 serotypes and their representative lineages were identified by different studies in this analysis, with DENV-2 being the most dominant. This agrees with findings from analysis of epidemics in all African sub-regions from 1960 to 2020 [2]. There were 21 study articles available for review using our search criteria. We observed a large difference in geographical distribution of DENV seroprevalence studies. Five geographical regions were identified in the DENV studies with 39 study sites [1,6-10,12,14,16,19,20,23,26-28,31-36]. There were higher infection rates among adults than in children [1,8,20,23,26,28]. This has been reported in other studies on DENV infection [17,37]. In children's studies, increasing age was found to be associated with overall DENV and other arbovirus infection, with outdoor activities and/or school attendance being associated with overall arbovirus seropositivity [20,31,32]. This could be due to vector exposure during outdoor excursions [17].
The fact that DENV infections have not been reported from most of the geographical regions in Kenya may not necessarily mean that those areas have no DENV seroprevalence. This could be due to their epidemiological situation being unknown. For instance, a recent study investigated the role of DENV infection as a cause of acute febrile disease in previously unreported non-epidemic settings in rural (Taita-Taveta County) and urban (Kibera slums, Nairobi) study areas in Kenya [23]. These areas were found to be seropositive. Detection of even one case in a population is a potential risk if the transmitting vectors are available [38]. This study has revealed unreported areas which are non-epidemic for consideration in national control strategies and surveillance to put in place preventative measures to avert outbreaks.
Our study has revealed a vast population across Kenya, especially rural settings, that has not had seroprevalence investigated. Of the 24 studies in our analysis, 4 were in rural sites [20,23,31,36], 6 were in both rural and urban sites [6,7,12,16,19,35], and the rest were in urban sites (Figure 2). While the task may appear overwhelming, it's possible to employ a systematic method inspired by a 2014 study conducted in Kenya [39] to pinpoint areas with the highest and lowest dengue risk. This approach utilizes a geospatial analysis similarity search to pinpoint these specific locations. It involves the creation of a risk map by merging environmental susceptibility analysis and geographic information systems (GIS), subsequently comparing dengue prevalence areas with all other regions. To enhance its reliability, the model also integrates bioclimatic factors, elevation data, and mosquito habitat information as inputs. This model can then be utilized to identify the areas with the highest risk, thus prioritizing them for further investigation.
Eight sporadic DENV outbreaks were reported during the ten-year study period, dominated with DENV-2 [9,13,14,27,33], but also DENV-3 and DENV-4 [27] and DENV-1[6]. During the outbreaks, the expression of the disease was relatively mild, although there were instances of hemorrhagic symptoms [9,14], and two unfortunate fatalities [13,14]. Mild disease manifestation could be due to the effective interventions which included distributing rapid dengue diagnostic kits to local health clinics and deploying response teams with the responsibility of raising community awareness about infection prevention. In addition, mosquito control activities were conducted, and health authorities carried out indoor and outdoor residual spraying [6]. To mitigate future outbreaks, such interventions should be strengthened.
Our study revealed that there is continued circulation of DENV infections across Kenya, with variable transmission patterns across different regions, but relatively homogenous within districts. In addition, DENV infections continue to circulate between epidemics or outbreaks (18 out of the 24 studies). This finding is supported by other studies on DENV infectivity [17,18]. There seems to be lack of definitive tests as accurate diagnosis available to fully appreciate the range and distribution of DENV infection in risk populations. Therefore, there is need for further development and use of techniques that can conclusively determine past exposures as well as low-level circulation of DENV. In addition, interaction with other arboviral infections (e.g., yellow fever, Rift Valley fever, chikungunya, o'nyong'nyong, West Nile virus, and zika virus) and parasitic infection (e.g., malaria) during inter-epidemic periods should be investigated [6,8,32]. This will enable appropriate control measures guided by accurate diagnosis targeting specific vectors responsible for the diseases.
Study designs can be categorized as passive facility-based fever surveillance, cost-of-illness surveys, serological surveys, and healthcare utilization surveys [21]. Our study revealed three main categories: health facility-based fever serosurveys (17), community-based serological surveys (4), and retrospective studies (3). Health facility-based fever serosurvey was the most preferred study design (17/24). This study further revealed that female gender was associated with overall DENV seropositivity in facility-based serosurveys [8,23,32]. This could be associated with differing healthcare seeking behavior where females have been reported elsewhere to seek more than males [40,41]. There is need for standardization of hospital-based surveillance and community-based serological designs in studying DENV infections. In addition, fever surveillance data should be robust, enabling calculation of several variables e.g., age-specific incidence rates, comparison of symptom presentations between DENV and non-DENV infections, multivariable logistic regression, seroprevalence, force of infection etc [21].
Studies in this paper diagnosed DENV using Enzyme immunoassays ELISA (IgA, IgG, and IgM) in combination with plaque reduction neutralization test (PRNT), ELISA- IgG and IgM in combination with Plaque Reduction Neutralization Tests (PRNTs), DENV IgM EIA, IgG IFA, ELISA- IgG solo, and ELISA- IgM solo. Other diagnostic tests included Viral RNA screening, Dengue NS1 antigen test, reverse transcription PCR (RT-PCR), quantitative real time reverse transcription PCR (qRT-PCR), real time reverse transcription PCR (rRT-PCR), a combination of Rapid Diagnostic Tests (RDT), RT-PCR and IgM/IgG ELISA, plaque reduction neutralization tests (PRNTs) solo, and sequencing, which included next generation sequencing (NGS). There is an inadequacy of detecting DENV solely by plaque reduction neutralization tests (PRNTs) because of possible cross-reactivity. Also, using solely anti-DENV IgM antibody responses in detecting DENV in individuals who have been vaccinated within 2 months will give false results [42]. A more reliable diagnosis for detecting DENV infection has been reported to be a combination of NS1 and IgM/IgG/IgA ELISAs, or ELISAs and PCR [19,27,42].
Accurate and efficient DENV diagnosis plays a crucial role in confirming cases. The virus can be detected during the viremic phase, typically within the first five days of illness, in serum, plasma, and peripheral blood mononuclear cells. Dengue NS1 antigen levels can be identified using enzyme-linked immunoassay (ELISA), with testing conducted through enzyme-immunoassays (EIAs) or immune-chromatographic (ICT) techniques. These assays are highly specific for differentiating between various flaviviruses. For early detection during the viremic phase, conventional and real time RT PCR, nested PCR, multiplex PCR, and nucleic acid sequence-based amplification (NASBA) are sensitive and relatively rapid methods. Additional tests include assessing anti-dengue specific IgM and IgG through ELISA [43]. However, these are sophisticated tests that are rare in resource-limited settings.
Improved diagnosis will enhance detection of underlying or co-circulating infections as well as detecting re-emerging and emerging disease threats caused by various pathogens associated with febrile illness manifestations in human populations [6,8-10,12,28,32]. This need has also been emphasized for the whole African continent [2]. The cost implications of definitive diagnosis can be prohibitive in resource poor facilities. This could be the reason why some studies conducted in patients with acute dengue used rapid diagnostic tests. Quick antigen tests, like the NS1 rapid diagnostic (RD) test, exhibit reasonable sensitivity, ranging from 71% to 80%, and high specificity (100%) when it comes to identifying dengue infection overall. However, they are unable to differentiate between serotypes [44]. Therefore, these results need to be scaled up with improved diagnostic techniques.
A cost-effective alternative for detecting dengue infection is the identification of anti-DENV IgM in urine, a highly specific marker for laboratory-confirmed dengue, as reported in one study [45]. This method holds promise for implementation in resource-constrained settings. In addition, recent research has explored the use of anti-aGal antibodies (IgM, IgG, IgG1, and IgG2) for diagnosing DENV. These antibodies target a specific glycan found in the virus, likely originating from mosquito salivary glands, which might play a role in the initial infection when DENV is introduced into the bloodstream through mosquito saliva [46]. Furthermore, a novel approach for DENV detection is the BioFire Global Fever Panel (GF Panel). This multiplex nucleic acid amplification test is performed directly on whole blood samples using the BioFire FilmArray System. It enables the diagnosis of various pathogens responsible for acute febrile illnesses, including DENV [47]. These diagnostic techniques could also be adopted in Kenya.
DENV-1-4 are graded into distinct genotypes; American (AM), Asian/American (AA), Asian I (AI), Asian II (AII), Cosmopolitan (C), and Sylvatic (S). In addition, new lineages have emerged from DENV-2 due to its ease of transmission and frequent occurrence, and the inevitable changing evolutionary pressure [26]. Most studies in this review did not provide sequencing, and consequently, genealogy results. Eleven out of the 24 studies did not do PCR or sequencing [1,16,20,23,31,32,34,36], with [16,23,31] having two studies each. Thus, the DENV genotype which was responsible for the infection will never be known. This supports reports that indicate that there is under-representation of Africa DENV sequences in databases [10,27,33]. The few genealogy studies in this paper indicated the Asian genotype was most isolated (5), African (3) and Cosmopolitan (3). In addition, there was one American (Brazil) genotype, but the Asian/American and Sylvatic genotypes were not revealed.
There is need to investigate regional variation in DENV strain predominance, as well as recognize the molecular differences between DENV strains that are detected during routine surveillance and those detected during outbreaks [26]. Furthermore, genealogy studies suggest an intercontinental connection in the spread of DENV. Hence, it is essential to consider both local and international strategies when formulating interventions to mitigate DENV infections. Information on circulating DENV serotypes in Kenya and their origin is important for vaccine development [7,10,27,33].
As there is currently no available vaccine to shield humans from DENV, the most successful approach to preventing DENV infections is to minimize interactions between people and the disease vectors. DENV suspected patients undergo diagnosis and positive cases are managed. Currently, Kenya employs integrated vector management (IVM) strategies to combat DENV infections. These measures involve entomological surveillance conducted by both governmental and non-governmental organizations in collaboration with the international community. They also include the application of synthetic pesticides, particularly pyrethroids, in indoor residual spraying (IRS), as well as the distribution of long-lasting insecticide nets (LLIN). Additionally, insect growth regulators (IGRs), potent insecticides derived from chemical or botanical sources, have been utilized [48].
In Europe, dengue vectors have been identified, but there is no active virus transmission. This is attributed to their risk management plans, which prioritize strategies for lowering the risk of transmission. These strategies encompass swift investigation of isolated cases, continuous monitoring of vector populations and their prevalence, social mobilization initiatives, and integrated vector management. In the event of a confirmed case, the response is elevated to an epidemic level to interrupt further transmission. Kenya should endeavor to implement these strategies to attain a low risk of DENV transmission [49,50]. Our study has identified Kenyan counties for which the DENV seroprevalence status remains unknown or unreported. This information is valuable for extending these interventions to previously unaddressed areas.
We cannot overemphasize the need for further research to accurately determine the burden and impact of DENV infection in different localities in Kenya. There is need for sufficient monitoring and increase in surveillance of these infections amongst patients presenting with fever in health facilities, as well as a better understanding of the ecology of arboviruses [6,23,36].
Limitations of the study
One of the limitations of this study, which may affect interpretation of our findings, was the underestimation of the number of DENV incidences and outbreaks in Kenya. This was occasioned by the fact that some of these incidences are reported in print media (which was not included in our selection criteria), and not in scientific published articles. Although no single mechanism for data collection is adequate to meet the needs for public health decision making, public health surveillance is essential and is reported to be one of the major mechanisms. However, most passive surveillance data are incomplete [51]. Our review was limited to scientific literature since one form of dissemination of surveillance data is peer-reviewed journals [52]. In addition, seroprevalence surveys provide more data. For instance, in our study, besides information on DENV infection and/or epidemic activity, information regarding the broad regional set-up, study site, period of study, study population, sample size, number of DENV-positive cases, case identification procedure, serotype and genotype was available for our review.
Other limitations included the retrospective nature of some of the reports, incomplete datasets, relatively small sample sizes, and inappropriate definition or selection of study population. Furthermore, different DENV identification procedures were used in the analyzed studies, which translates into different quality of the used serologic assays or molecular techniques. This posed a risk in methodological bias during the quality assessment of included studies. Nonetheless, the overall methodological risk bias of our study was low (3) (Table 2). Thus, we are confident that our findings provide useful information for public policy decisions and clinical practice.
This study on DENV infection and epidemics in Kenya in the period 2010 to 2020 has revealed surveillance gaps which will guide strategies for reducing the burden of the disease. We provide evidence for serosurveys in five geographical regions and eleven counties in the country. Coast is the most studied region with Mombasa being the most sampled site. Thirty six (36) out of the 47 (76.6%) Kenyan counties' DENV seroprevalence status is unknown or unreported. Our study has also revealed that DENV infection is unrecognized and/or unreported in most areas of the country, especially rural Kenya. In addition, it reveals that there is transmission between epidemics in the reported areas.
Recommendations
We recommend robust surveillance of DENV infections and strict implementation measures in Kenya to improve human public health and reduce the possibility of potential epidemics. The public health and vector control strategies designed to mitigate the burden of dengue in Kenya should consider the unreported and non-epidemic areas in the country as high priority areas because they could be potential “breeding grounds” for DENV outbreaks. In addition, more data is needed to document clinical and epidemiologic patterns of dengue in Kenya as well as genealogy of circulating genotypes for vaccine development. In addition, knowledge of spatial-temporal dynamics of dengue circulation is critically needed for better response in control strategy [12]. Control efforts might be more effective if targeted at a district level, as opposed to country-wide, which would be too broad or village-specific, which might be too narrow [17]. For an effective study, this gap can be addressed by adopting population-based dengue surveillance and seroprevalence studies [21].
Finally, there is need to adopt best practices and newer approaches in controlling infectious diseases. One such proposal is a “One Health” concept, which entails creation of a multidisciplinary platform consisting of a team of members with a range of expertise to monitor, respond to and prevent major outbreaks [53]. The team could consist of agriculturalists, animal husbandry specialists, ecologists, physicians, public health officers, researchers, climatologists, vector biologists, veterinarians, and viral phylogeneticists. Their concerted co-operation and collaboration will unravel the relationship and interconnectedness between animals, the environment, and humans. This is because the health and wellbeing of humans is covertly tied to the health of animals and their environment, and the other way round.
Therefore, a national surveillance program should plan for serosurveys in the unreported areas with proper representativeness of the population to investigate seroprevalence of DENV infections. This will provide a powerful tool to better understand and plan for the response to dengue outbreaks in areas with limited surveillance representation. Together with adoption of new and best practices such as the East African Community (EAC) intergovernmental body regional network for rapid response to epidemics, and the “One Health” multidisciplinary platform, DENV infections in Kenya will be under control.
What is known about this topic
- DENV is endemic in Kenya, with all the four serotypes (DENV- 1-4);
- DENV is a major public health problem in Kenya due to frequent outbreaks.
What this study adds
- This review revealed a surveillance gap in dengue infection surveys in Kenya, with DENV infection being un-recognized and/or unreported in 76.6% of the country;
- It showed a methodological bias in DENV studies which affect quality of data generated, thus, need for harmonization of future study procedures;
- It further revealed that DENV-2 is the dominant circulating serotype and circulating DENV genealogies have an intercontinental link, which information is important for vaccine development as well as local and international intervention strategies.
The authors declare no competing of interests.
Carren Bosire conceptualized the review. Carren Bosire and Kavinya Mwendwa evaluated quality of selected DENV infection seroprevalence studies. Francis Mutuku, Bryson Ndenga, Makenzi Nzaro, Kavinya Mwendwa and Angelle Desiree LaBeaud critically reviewed, and co-authored the manuscript. All authors have read and agreed to the final manuscript.
This research was supported by library resources and funding from the Office of Registrar, Research, Partnership and Extension of Technical University of Mombasa.
Table 1: eligibility criteria
Table 2: risk of bias scores for individual studies included in the review
Table 3: DENV positive samples
Table 4: reported DENV outbreaks in Kenya (2010-2020)
Table 5: DENV identification procedures
Table 6: serotypes and lineages of DENV infections in Kenya (2010-2020)
Figure 1: PRISMA flow chart illustrating screening and filtering of articles for eligibility
Figure 2: map of Kenya showing regional distribution of DENV seroprevalence study sites (2010-2020)
- Munyuga K, Ng'ang'a J, Inoue S, Syengo KC. Co-Circulation Evidence of Dengue Virus Serotypes at the Kenyan Coast in 2014, 2015. IOSR J Pharm Biol Sci. Dec 2016;11(6):83-87.
- Mwanyika GO, Mboera LEG, Rugarabamu S, Ngingo B, Sindato C, Lutwama JJ, et al. Dengue Virus Infection and Associated Risk Factors in Africa: A Systematic Review and Meta-Analysis. Viruses. Mar 2021;13(4):536. PubMed | Google Scholar
- World Health Organisation. Dengue and severe dengue. Accessed May 19, 2023.
- Bhatt S, Gething PW, Brady OJ, Messina JP, Farlow AW, Moyes CL, et al. The global distribution and burden of dengue. Nature. Apr 2013;496(7446):504-507. PubMed | Google Scholar
- Brady OJ, Gething PW, Bhatt S, Messina JP, Brownstein JS, Hoen AG, et al. Refining the global spatial limits of dengue virus transmission by evidence-based consensus. PLoS Negl Trop Dis. 2012;6(8):e1760. PubMed | Google Scholar
- Konongoi L, Ofula V, Nyunja A, Owaka S, Koka H, Makio A, et al. Detection of dengue virus serotypes 1, 2 and 3 in selected regions of Kenya: 2011-2014. Virol J. 2016 Nov 4;13(1):182. PubMed | Google Scholar
- Langat SK, Eyase FL, Berry IM, Nyunja A, Bulimo W, Owaka S, et al. Origin and evolution of dengue virus type 2 causing outbreaks in Kenya: Evidence of circulation of two cosmopolitan genotype lineages. Virus Evol. 2020 Jun 2;6(1):veaa026. PubMed | Google Scholar
- Ngoi CN, Price MA, Fields B, Bonventure J, Ochieng C, Mwashigadi G, et al. Dengue and Chikungunya Virus Infections among Young Febrile Adults Evaluated for Acute HIV-1 Infection in Coastal Kenya. PLoS One. 2016 Dec 12;11(12):e0167508. PubMed | Google Scholar
- Obonyo M, Fidhow A, Ofula V. Investigation of laboratory confirmed Dengue outbreak in North-eastern Kenya, 2011. PLoS One. 2018 Jun 7;13(6):e0198556. PubMed | Google Scholar
- Pollett S, Gathii K, Figueroa K, Rutvisuttinunt W, Srikanth A, Nyataya J, et al. The evolution of dengue-2 viruses in Malindi, Kenya and greater East Africa: Epidemiological and immunological implications. Infect Genet Evol. Jun 2021;90:104617. PubMed | Google Scholar
- Bhatia S, Bansal D, Patil S, Pandya S, Ilyas QM, Imran S. A Retrospective Study of Climate Change Affecting Dengue: Evidences, Challenges and Future Directions. Front Public Health. May 2022;10:884645. PubMed | Google Scholar
- Shah MM, Ndenga BA, Mutuku FM, Vu DM, Grossi-Soyster EN, Okuta V, et al. High Dengue Burden and Circulation of 4 Virus Serotypes among Children with Undifferentiated Fever, Kenya, 2014-2017. Emerg Infect Dis. Nov 2020;26(11):2638-2650. PubMed | Google Scholar
- World Health Organization. African Region. Outbreaks and Emergencies Bulletin, Week 28: 8 - 14 July 2017. Accessed May 19, 2023.
- Ellis EM, Neatherlin JC, Delorey M, Ochieng M, Mohamed AH, Mogeni DO, et al. A Household Serosurvey to Estimate the Magnitude of a Dengue Outbreak in Mombasa, Kenya, 2013. PLoS Negl Trop Dis. Apr 2015;9:e0003733. PubMed | Google Scholar
- Chan KR, Ismail AA, Thergarajan G, Raju CS, Yam HC, Rishya M, et al. Serological cross-reactivity among common flaviviruses. Front Cell Infect Microbiol. 2022;12:975398. PubMed | Google Scholar
- Hortion J, Mutuku FM, Eyherabide AL, Vu DM, Boothroyd DB, Grossi-Soyster EN, et al. Acute Flavivirus and Alphavirus Infections among Children in Two Different Areas of Kenya, 2015. Am J Trop Med Hyg. 2019 Jan;100(1):170-173. PubMed | Google Scholar
- Mease LE, Coldren RL, Musila LA, Prosser T, Ogolla F, Ofula VO, et al. Seroprevalence and distribution of arboviral infections among rural Kenyan adults: A cross-sectional study. Virol J. Jul 2011;8:371. PubMed | Google Scholar
- Sutherland LJ, Cash AA, Huang YJ, Sang RC, Malhotra I, Moormann AM, et al. Serologic evidence of arboviral infections among humans in Kenya. Am J Trop Med Hyg. 2011 Jul;85(1):158-61. PubMed | Google Scholar
- Vu DM, Mutai N, Heath CJ, Vulule JM, Mutuku FM, Ndenga BA, et al. Unrecognized Dengue Virus Infections in Children, Western Kenya, 2014-2015. Emerg Infect Dis. 2017 Nov;23(11):1915-1917. PubMed | Google Scholar
- Grossi-Soyster EN, Cook EAJ, de Glanville WA, Thomas LF, Krystosik AR, Lee J, et al. Serological and spatial analysis of alphavirus and flavivirus prevalence and risk factors in a rural community in western Kenya. PLoS Negl Trop Dis. Oct 2017;11(10):e0005998. PubMed | Google Scholar
- Lim JK, Carabali M, Lee JS, Lee KS, Namkung S, Lim SK, et al. Evaluating dengue burden in Africa in passive fever surveillance and seroprevalence studies: protocol of field studies of the Dengue Vaccine Initiative. BMJ Open. 2018 Jan 21;8(1):e017673. PubMed
- Gainor EM, Harris E, LaBeaud AD. Uncovering the Burden of Dengue in Africa: Considerations on Magnitude, Misdiagnosis, and Ancestry. Viruses. 2022 Jan 25;14(2):233. PubMed | Google Scholar
- Masika MM, Korhonen EM, Smura T, Uusitalo R, Vapalahti K, Mwaengo D, et al. Detection of dengue virus type 2 of Indian origin in acute febrile patients in rural Kenya. PLoS Negl Trop Dis. Mar 2020;14(3):e0008099. PubMed | Google Scholar
- Johnson BK, Ocheng D, Gichogo A, Okiro M, Libondo D, Kinyanjui P, et al. Epidemic dengue fever caused by dengue type 2 virus in Kenya: preliminary results of human virological and serological studies. East Afr Med J. Dec 1982;59(12):781-784. PubMed
- Sang RC, Dunster LM. The growing threat of arbovirus transmission and outbreaks in Kenya: a review. East Afr Med J. 2001 Dec;78(12):655-61. PubMed | Google Scholar
- Gathii K, Nyataya JN, Mutai BK, Awinda G, Waitumbi JN. Complete Coding Sequences of Dengue Virus Type 2 Strains from Febrile Patients Seen in Malindi District Hospital, Kenya, during the 2017 Dengue Fever Outbreak. Genome Announc. 2018 Apr 12;6(15):e00076-18. PubMed | Google Scholar
- Muthanje EM, Kimita G, Nyataya J, Njue W, Mulili C, Mugweru J, et al. March 2019 dengue fever outbreak at the Kenyan south coast involving dengue virus serotype 3, genotypes III and V. PLOS Glob Public Health. 2022 Mar 24;2(3):e0000122. PubMed | Google Scholar
- Lim JK, Matendechero SH, Alexander N, Lee JS, Lee KS, Namkung S, et al. Clinical and epidemiologic characteristics associated with dengue fever in Mombasa, Kenya. Int J Infect Dis. 2020 Nov;100:207-215. PubMed | Google Scholar
- Liberati A, Altman DG, Tetzlaff J, Mulrow C, Gøtzsche PC, Ioannidis JP, et al. The PRISMA statement for reporting systematic reviews and meta-analyses of studies that evaluate health care interventions: explanation and elaboration. J Clin Epidemiol. 2009 Oct;62(10):e1-34. PubMed | Google Scholar
- Hoy D, Brooks P, Woolf A, Blyth F, March L, Bain C, et al. Assessing risk of bias in prevalence studies: modification of an existing tool and evidence of interrater agreement. J Clin Epidemiol. Sep 2012;65(9):934-9. PubMed | Google Scholar
- Awando JA, Ongus JR, Ouma C, Mwau M. Seroprevalence of Anti-Dengue Virus 2 Serocomplex Antibodies in out-patients with fever visiting selected hospitals in rural parts of Western Kenya in 2010-2011: a cross sectional study. Pan Afr Med J. Oct 2013;16:73. Google Scholar
- Inziani M, Adungo F, Awando J, Kihoro R, Inoue S, Morita K, et al. Seroprevalence of yellow fever, dengue, West Nile and chikungunya viruses in children in Teso South Sub-County, Western Kenya. Int J Infect Dis. 2020 Feb;91:104-110. PubMed | Google Scholar
- Kamau E, Agoti CN, Ngoi JM, de Laurent ZR, Gitonga J, Cotten M, et al. Complete Genome Sequences of Dengue Virus Type 2 Strains from Kilifi, Kenya. Microbiol Resour Announc. 2019 Jan 24;8(4):e01566-18. PubMed | Google Scholar
- Nzou S, Mwau M. A health facility based serosurvey of circulating Dengue in Trans nzoia, Western Kenya. Int J Infect Dis. Dec 2016;53:159. Google Scholar
- Vu DM, Ripp K, Mutai N, Ndenga BA, Heath C, LaBeaud AD. Dengue virus and malaria co-infection in Kenyan children. Annals of Global Health. 2016 Aug 20;82(3):435-6. PubMed | Google Scholar
- Vu DM, Banda T, Teng CY, Heimbaugh C, Muchiri EM, Mungai PL, et al. Dengue and West Nile Virus Transmission in Children and Adults in Coastal Kenya. Am J Trop Med Hyg. 2017 Jan 11;96(1):141-143. PubMed | Google Scholar
- Stanaway JD, Shepard DS, Undurraga EA, Halasa YA, Coffeng LE, Brady OJ, et al. The global burden of dengue: an analysis from the Global Burden of Disease Study 2013. Lancet Infect Dis. 2016 Jun;16(6):712-723. PubMed | Google Scholar
- Chepkorir E, Tchouassi DP, Konongoi SL, Lutomiah J, Tigoi C, Irura Z, et al. Serological evidence of Flavivirus circulation in human populations in Northern Kenya: an assessment of disease risk 2016-2017. Virol J. 2019 May 17;16(1):65. PubMed | Google Scholar
- Attaway DF, Jacobsen KH, Falconer A, Manca G, Rosenshein Bennett L, Waters NM. Mosquito habitat and dengue risk potential in Kenya: alternative methods to traditional risk mapping techniques. Geospat Health. 2014 Nov;9(1):119-30. PubMed | Google Scholar
- Bertakis KD, Azari R, Helms LJ, Callahan EJ, Robbins JA. Gender differences in the utilization of health care services. J Fam Pract. 2000 Feb;49(2):147-52. PubMed | Google Scholar
- Yeatman S, Chamberlin S, Dovel K. Women's (health) work: A population-based, cross-sectional study of gender differences in time spent seeking health care in Malawi. PLoS One. 2018 Dec 21;13(12):e0209586. PubMed | Google Scholar
- Hunsperger EA, Duarte Dos Santos CN, Vu HTQ, Yoksan S, Deubel V. Rapid and accurate interpretation of dengue diagnostics in the context of dengue vaccination implementation: Viewpoints and guidelines issued from an experts group consultation. PLoS Negl Trop Dis. 2017 Sep 7;11(9):e0005719. PubMed | Google Scholar
- Bhat VG, Chavan P, Ojha S, Nair PK. Challenges in the Laboratory Diagnosis and Management of Dengue Infections. Open Microbiol J. 2015 Jul 31;9:33-7. PubMed | Google Scholar
- Santoso MS, Yohan B, Denis D, Hayati RF, Haryanto S, Trianty L, et al. Diagnostic accuracy of 5 different brands of dengue virus non-structural protein 1 (NS1) antigen rapid diagnostic tests (RDT) in Indonesia. Diagn Microbiol Infect Dis. 2020 Oct;98(2):115116. PubMed | Google Scholar
- Caraballo E, Poole-Smith BK, Tomashek KM, Torres-Velasquez B, Alvarado LI, Lorenzi OD, et al. The detection of anti-dengue virus IgM in urine in participants enrolled in an acute febrile illness study in Puerto Rico. PLoS Negl Trop Dis. 2020 Jan 29;14(1):e0007971. PubMed | Google Scholar
- Olajiga OM, Maldonado-Ruiz LP, Fatehi S, Cardenas JC, Gonzalez MU, Gutierrez-Silva LY, et al. Association of dengue infection with anti-alpha-gal antibodies, IgM, IgG, IgG1, and IgG2. Front Immunol. 2022 Oct 14;13:1021016. PubMed | Google Scholar
- Manabe YC, Betz J, Jackson O, Asoala V, Bazan I, Blair PW, et al. Clinical evaluation of the BioFire Global Fever Panel for the identification of malaria, leptospirosis, chikungunya, and dengue from whole blood: a prospective, multicentre, cross-sectional diagnostic accuracy study. Lancet Infect Dis. 2022 Sep;22(9):1356-1364. PubMed | Google Scholar
- Karungu S, Atoni E, Ogalo J, Mwaliko C, Agwanda B, Yuan Z, et al. Mosquitoes of Etiological Concern in Kenya and Possible Control Strategies. Insects. 2019 Jun 16;10(6):173. PubMed | Google Scholar
- Schaffner F, Mathis A. Dengue and dengue vectors in the WHO European region: past, present, and scenarios for the future. Lancet Infect Dis. 2014 Dec;14(12):1271-80. PubMed | Google Scholar
- Altmetric. Global strategy for dengue prevention and control 2012-2020. Accessed Sept 21, 2023.
- Stansfield SK, Walsh J, Prata N, Evans T. Information to Improve Decision Making for Health. In: Jamison DT, Breman JG, Measham AR, et al., editors. Disease Control Priorities in Developing Countries. 2nd edition. Washington (DC): The International Bank for Reconstruction and Development/The World Bank; 2006. Chapter 54. Co-published by Oxford University Press, New York.
- Murray J, Cohen AL. Infectious Disease Surveillance. Int Encycl Public Health. 2017;222-229. PubMed
- Zumla A, Dar O, Kock R, Muturi M, Ntoumi F, Kaleebu P, et al. Taking forward a 'One Health' approach for turning the tide against the Middle East respiratory syndrome coronavirus and other zoonotic pathogens with epidemic potential. Int J Infect Dis. 2016 Jun;47:5-9. PubMed | Google Scholar