Bacterial ecology and antibiotic susceptibility profile of isolated strains from surfaces and medical devices in some departments of the Jordan Medical Services, Cameroon: a descriptive cross-sectional study
Nadege Lylian Doumou AMvomo, Cybelle Kemayou Kabit Sama, Rodrigue Bouopda, Flora Christelle Nkwinkwa Minkeza, Laure Patricia Mbopda, Cindy Loïda Yonga Tchoumi, Lesly Ornella Tchouangeum Kenmoe, Gilles Tchouassi Simeni, Eric Chokote Tolo, Ruth Ngongang, Esther Voundi Voundi
Corresponding author: Nadege Lylian Doumou AMvomo, Jordan Medical Services, Medical and Technical Department, Yaoundé, Cameroon 
Received: 13 Dec 2022 - Accepted: 19 May 2023 - Published: 24 May 2023
Domain: Bacteriology,Epidemiology
Keywords: Bacterial ecology, antibiotic, susceptibility profile, strains isolated, Jordan Medical services
©Nadege Lylian Doumou AMvomo et al. PAMJ-One Health (ISSN: 2707-2800). This is an Open Access article distributed under the terms of the Creative Commons Attribution International 4.0 License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Cite this article: Nadege Lylian Doumou AMvomo et al. Bacterial ecology and antibiotic susceptibility profile of isolated strains from surfaces and medical devices in some departments of the Jordan Medical Services, Cameroon: a descriptive cross-sectional study. PAMJ-One Health. 2023;11:6. [doi: 10.11604/pamj-oh.2023.11.6.38513]
Available online at: https://www.one-health.panafrican-med-journal.com/content/article/11/6/full
Research 
Bacterial ecology and antibiotic susceptibility profile of isolated strains from surfaces and medical devices in some departments of the Jordan Medical Services, Cameroon: a descriptive cross-sectional study
Bacterial ecology and antibiotic susceptibility profile of isolated strains from surfaces and medical devices in some departments of the Jordan Medical Services, Cameroon: a descriptive cross-sectional study
Nadege Lylian Doumou AMvomo1,2,&, Cybelle Kemayou Kabit Sama3, Rodrigue Bouopda3,4, Flora Christelle Nkwinkwa Minkeza1, Laure Patricia Mbopda1, Cindy Loïda Yonga Tchoumi1, Lesly Ornella Tchouangeum Kenmoe1, Gilles Tchouassi Simeni1, ![]() Eric Chokote Tolo1,2, Ruth Ngongang1,2, Esther Voundi Voundi
Eric Chokote Tolo1,2, Ruth Ngongang1,2, Esther Voundi Voundi
&Corresponding author
Introduction: within the framework of microbiological surveillance of the hospital environment, the present study aims to identify the different bacterial species prevalent on hospital surfaces and devices in the various departments of the Jourdan Medical Services and to determine their antibiotic sensitivity profile
Methods: we conducted a descriptive cross-sectional study during May 2022, in different departments of the Jourdan Medical Services. Swabbing of surfaces and medical devices was performed and antibiotic susceptibility was determined by the Kirby-Bauer disc diffusion method, in accordance with the Antibiotic Committee of the French Society of Microbiology. The data thus collected were analyzed using Excel 2006.
Results: of 156 samples, 86 were culture positive and 100 bacteria were isolated. Klebsiella pneumoniae (17%), Enterobacter cloacae (17%), Staphylococcus aureus (16%) and Escherichia coli (11%) were mainly isolated. The most soiled surfaces were trolleys (12%), taps (12%), door handles (11%) and tables (10%). For medical devices, the most soiled were injectomats (3%), thermometers (3%), and blood pressure cuffs (3%). Seratia liquefaciens, Enterobacter cloacae, Kblesiella pneumoniae and Escherichia coli produced Extended Spectrum Betalactamase (23.3%). Of the Staphylococcus aureus isolates, 81.65% (13/16) were Methicillin-Resistant strains.
Conclusion: the microbial ecology of surfaces and medical devices is dense and consists of multi-resistant bacteria. Cleaning and disinfection should be regularly evaluated through monitoring programs in hospital departments.
Healthcare-associated infections (HAIs) are a major public health problem, particularly as they contribute to the global threat of antibiotic resistance, which is expected to cause 10 million deaths per year by 2050 and will be the leading cause of death in the world [1]. Surfaces and medical devices are colonized by microorganisms in hospitals, which can come from patients, the air, visitors and nursing staff [2]. Health care institutions are a favorable ecosystem for multi-resistant bacteria for two reasons: 25% of hospitalized patients receive antibiotics on a permanent basis, with a selection pressure of germs, and the very easy human-to-human transmission of bacteria due to the promiscuity and density of care and nursing [1]. This constitutes a risk factor for the occurrence of nosocomial infections, a real threat, both to the already precarious health of patients and to that of nursing staff and visitors. These infections increase healthcare costs and length of stay and are the leading cause of mortality and morbidity among hospitalized patients [3]. According to the World Health Organization (WHO), the impact of nosocomial infections and antimicrobial resistance on the lives of those affected is incalculable. More than 24% of patients with hospital-acquired sepsis and 52.3% of patients in intensive care units die each year. Deaths are doubled or tripled if patients have antimicrobial-resistant infections [4]. The micro-organisms involved in the hospital environment are most often multi-resistant to antibiotics, and the main source of diffusion of highly pathogenic strains in the departments [2]. The fight against nosocomial infections, and in particular those linked to germs in the hospital environment, is one of the priorities of health care institutions. Thus, the control of contamination of hospital surfaces requires microbiological controls [5]. A better understanding of the spread of bacteria responsible for nosocomial infections on surfaces and medical devices therefore appears to be a major area for research. Within the framework of microbiological surveillance of the hospital environment, the present study aims to identify the different bacterial species prevalent on hospital surfaces and devices in the various departments of the Jourdan Medical Services and to determine their antibiotic sensitivity profile. Our specific hypothesis is therefore that surfaces and medical devices are contaminated with antibiotic-resistant bacteria.
Study design: bacterial ecology, antibiotic sensitivity profile, bacteria, surfaces and medical devices.
Setting: we conducted a descriptive cross-sectional study from May 15th to July 5th, 2022 in the following 6 departments: Emergency and Hospitalization, Operating Room, Intensive Care Unit, Maternity Ward, Medical Analysis Laboratory and the waiting room of the Jourdan Medical Centre. This is a private hospital located in the Centre Cameroon Region, whose mission is to promote health and ensure the well-being of the population. The hospital has a capacity of 36 beds and houses all major medical specialties.
Participants: they were included in our study, hospital surfaces and medical devices found in the above six departments using swabs pre-moistened with sterile salt wate, in accordance with ISO/DIS 14698-1 [6]. They were chosen based on the high-risk infection they could cause must, especially to patients. The choice of sampling points was also based on the hygiene guide for environmental surfaces [7]. The method was non-probabilistic by convenience.
Variables: bacterial ecology and antibiotic susceptibility profile of isolated strains.
Bias: in order to avoid potentials bias, we carried out several controls on the culture media, antibiotics, equipments and the reagents.
Study size: of the 6 sites selected, one hundred and fifty-six (156) samples were taken from medical devices (n=39) and surfaces (n=117). The various samples were taken from cupboards (8), the operating field (1), chairs (2), trolleys (12), switches (15), gynecological beds (2), mattresses (3) benches (3), door handles (21), gallows (12), bed curtain (7), curtains (5), taps (12), tables (11), remote controls (3), anesthesia machine (2), hoovers (2), autoclave (1), bedpan (2); electric scalpel (1), surgical boxes (2), blood pressure cuff (3), wheelchairs (3), incubator (1), gown (2), incubators (2), injectomats (3), tape measure (2), microscopes (2), oximeter (2), poupinel (1), light strip (1), refrigerator (1), respirator (1), stethoscope (1), thermometer (4).
Quantitative variables: samples were taken in the early morning hours, using sterile swabs, pre-moistened with sterile salt water, in accordance with ISO/DIS 14698-1 [6], from surfaces and medical devices present in the wards during the study period. The technique consisted of moistening a swab with 0.9% sodium chloride solution and then passing it over the surface to be sampled, making parallel, close striations while turning the wet swab slightly. Sampling of the same area was repeated by making striations perpendicular to the first. The various swabs were placed in labelled cryotubes containing heart and brain broth and then quickly sent to the laboratory for bacteriological examination.
Isolation and phenotypic identification of aerobic bacteria: after transporting the samples to the laboratory, the cryotubes were incubated at 37�°C±2�°C for 18-24 hours. Sub culturing was systematically done on Columbia agar on the second day, followed by a new incubation at 37�°C±2�°C for 18-24 hours. On the third day, after description of the colonies if culture positive and GRAM staining, sub culturing was done in specific media, according to the GRAM results followed by a further incubation. Biochemical testing of bacterial isolates was performed using the API 20E gallery and the identification algorithm for Gram-positive cocci, Bacillus spp and non-fermenting Enterobacteriaceae.
Antibiogram of isolated bacteria: antibiotic susceptibility was tested by the Muller-Hinton agar disk diffusion technique [8]. From a visible culture, we made a bacterial suspension in salt solution to achieve a turbidity equivalent to that of the 0.5 standard of the McFarland range. Twenty two antibiotic discs (OXOID) were tested: Amoxicillin (20�μg), Amoxicillin-clavulanic acid (20�μg -10�μg), Piperacillin (30�μg), Piperacillin-tazobactam (30-6�μg), Ticarcillin (75�μg), Cefoxitin (30�μg), Ceftazidime (10�μg), Ceftriaxone (30�μg), Cefepime (30�μg), Imipenem (10�μg), Ciprofloxacin (30�μg), Levofloxacin (5�μg), Gentamycin (10�μg), Erythromycin (15�μg), Kanamycin (30�μg), Tobramycin (10�μg), Tetracycline (30�μg), Fusidic acid (10�μg), Rifampicin (5�μg), Chloramphenicol (30�μg), Vancomycin (30�μg) and Teicoplanin (30�μg). The choice of antibiotic discs was based on the Antibiotic Committee of the French Society of Microbiology 2021 [8]. The discs were placed firmly on the surface of the dry inoculated agar and the media were incubated at 37�°C±2�°C for 18-24 hours. The results of the susceptibility tests were interpreted according to the recommendations of the Antibiogram Committee of the French Society of Microbiology [8]. Phenotypic detection of extended-spectrum beta-lactamase (ESBL) in Enterobacteriaceae isolates was performed in vitro by the disc diffusion method also known as the double disc synergy test (a disc matching test) combining amoxicillin-clavulanic acid with a third generation cephalosporin. The appearance of a synergistic image between these antibiotics (champagne cork) indicates ESBL production by the strain [8].
Staphylococcus aureus: strains were tested for methicillin-resistance using a cefoxitime disc as part of a standard susceptibility test. Strains with an inhibition diameter of less than 22 mm were considered Methicillin-resistant Staphylococcus aureus (MRSA) [8].
Statistical methods: the results included the location of isolation, the germs identified and their antibiotic susceptibility. Descriptive analysis of the data was performed using Excel 2006.
Ethical considerations: this study was conducted in accordance with the ethical guidelines for research in Cameroon. We obtained ethical clearance from the School of Health Sciences of the Catholic University of Central Africa, N°2022/022178/CEIRSH/ESS/MBV.
Level of contamination by department: of the 156 samples taken, 86 showed a positive culture. Of these, 76.7% (66/86) surfaces and 23.3% (20/86) medical devices showed positive cultures, with a monomicrobial growth of 83.7% (72/86) and a polymicrobial growth of 16.3% (14/86) (Table 1).
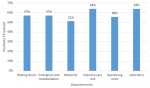
Positivity threshold per unit: from a bacteriological point of view, the positivity threshold per unit was 40.9% (9/22) in the Laboratory and Medical Analysis Unit, 64% (16/25) in the Operating Theatre Unit, 55.6% (15/27) in the Intensive Care Unit, 64% (16/25) in the Maternity Unit, 51.2% (22/43) in the Emergency and Hospitalization Unit; and in the waiting room 57.1% (8/14) (Figure 1).
Distribution of bacteria isolated per unit: Klebsiella pneumoniae, Enterobacter cloacae, Staphylococcus aureus and Escherichia coli were the main germs frequently isolated with a frequency of 17%, 17%, 16% and 11% respectively. The Emergency and Hospitalization department showed a high rate of isolated bacteria with a predominance of Escherichia coli (36%) (Table 2).
Distribution of bacteria isolated by surface: the most soiled surfaces were trolleys (12%), taps (12%), door handles (11%), tables (10%) and brackets (9%). Medical devices included injectomats (3%), thermometers (3%), blood pressure cuffs (3%), microscopes and aspirators (2% respectively) (Table 3).
Antibiotic susceptibility profile: in Staphylococaceae, we noted 100% resistance to Levofloxacin, Erythromycin, Kanamycin, Tobramycin and Tetracyclin; 93.3% to Ciprofloxacin and Gentamycin; 86.7% to Fusidic Acid and Chloramphenicol; and 80% to Cefoxitin. Among the Staphylococcus aureus isolated (13/16), strains resistant to Meticillin were noted (Meti-R = 81.65%). The isolated Enterobacteriaceae were 100% resistant to Ceftazidime, 96.7% to Piperacillin and Piperacillin + Tazobactam, 93.3% to Amoxicillin, 88.3% to Gentamycin, 86.7% to Ceftriaxone, 81.7% to Amoxicillin + Clavulanic Acid, 80% to Cefoxitin and 76.7% to Cefepime. Of the 60 enterobacteria isolated, 14 produced an Extended Spectrum Betalactamase (EBLSE =23.3%), including Serratia liquefaciens (1/1), Enterobacter cloacae (4/26), Klebsiella pneumonia (3/20) and Escherichia coli (6/11). The genus Bacillus showed 100% resistance to all antibiotics tested. The genus Pseudomonas showed 100% resistance to the antibiotics tested with the exception of Gentamycin (66.7%) (Table 4).
Surfaces and medical devices are colonized by microorganisms in the hospital environment, which may originate from patients, the air, visitors and nursing staff [2]. Microbiological sampling of the environment in hospital wards makes it possible to determine the microbial reservoir which is the origin of hospital-acquired infections [2]. These microbial reservoirs are one of the key indicators of poor hospital hygiene [7]. In our study, the overall bacterial contamination rate of surfaces and medical devices was 55.1%. This rate is close to the prevalence rate of 44.54% obtained from the Surfaces and Medical Devices in the Intensive Care Unit of the University Hospital Centre (CHU) of Treichville [2]; and to the studies carried out in Nigeria and Ethiopia which reported lower positivity rates, i.e. 46.7% and 39.6% respectively [9]. However, this rate is relatively low compared to work conducted at the Provincial Hospital Centre (PHC) of Mohammedia, Morocco where a rate of 88.4% was obtained [9]. The most contaminated units were the operating room (64%), the maternity ward (64%), the intensive care unit (55.6%), the waiting room (57.1%) and the emergency room (51.2%). These results are similar to the work carried out at the PHC where the most contaminated services were the maternity ward (92.5%), the Emergency ward (87.5%) and the operating theatre (75%); and to the work carried out at the Treichville University Hospital where the Intensive care unit was the most contaminated (44.5%). This difference can be explained by the fact that contamination varies both qualitatively and quantitatively over time, from one establishment to another and, within the same establishment, according to the services, patients, care and techniques used [9]. On the other hand, it could be due to the existence of more efficient equipment in European and some African bacteriology laboratories, contrary to our limited working conditions. The high level of contamination in the waiting room (57.1%) could be explained by the constant presence of nursing staff (doctors, nurses) during seminars and postgraduate courses.
The most contaminated surfaces were trolleys, gallows, taps and door handles. The most contaminated medical devices were Injectomats, blood pressure cuffs, hoovers and microscopes. Our results are similar to the work carried out at the Treichville University Hospital, where the sink and cupboards were the most contaminated surfaces, followed by injectomats and blood pressure cuffs [2]; and to the work carried out at the Yopougon University Hospital, Abidjan, Ivory Coast, where the most contaminated sites were the taps and door handles [10]. The high bacterial density on almost all surfaces could be the result of a lack of cleaning or the use of ineffective antiseptics, with the consequence that other surfaces are contaminated by pathogens. Bacteriologically, several species were isolated from contaminated surfaces and medical devices. The most prevalent were Staphylococcus aureus (19.2%), Klebsiella pneumoniae (17.8%), Escherichia coli (15.1%), Enterobacter cloacae (15.1%) and Bacillus spp. These results are not far from those found in Morocco where proportions of 27%, 20%, 16% and 5% were recorded for Bacillus spp, S. aureus, K. pneumoniae, P. aeruginosa and E. cloacae respectively [9]. Staphylococcus aureus, Klebsiella pneumoniae and Pseudomonas aeruginosa were the main germs frequently isolated, due to the fact that these bacteria are very often involved in severe infections, in addition to their nosocomial nature [2].
P. aeruginosa is responsible for severe nosocomial pathologies that can reach, for example, 70% of lethality in the case of nosocomial pneumopathy. The severity of these infections is linked to the virulence of these bacteria but also to their occurrence in debilitated conditions with a significant favoring effect of invasive devices leading to hospitalization [11]. They are ubiquitous aerobic bacteria, widely present in the environment, growing between 4 and 41°C. They usually survive only a few hours in a dry environment. However, they can survive and multiply in the water environment even in the absence of nutrients, especially in hospitals [11]. The high percentage of bacterial strains is related to poor hospital hygiene. The presence of Enterobacter Cloacae and Escherichia coli, in this work, testifies to fecal contamination [2]; and this confirms this poor hygiene in the care units; which constitutes a determining factor in the risk of nosocomial infections. Indeed, the significant colonization of various materials and surfaces constitutes a real risk of the transmission of resistant bacteria, which can be the cause of severe nosocomial infections [2]. The Bacillus genus is made up of spore-forming and telluric bacteria that are ubiquitous and found in soil, water, dust, plants and human and animal feces. A rate of 7% was isolated in this study which is comparable to the work carried out at the PHC in Morocco where a rate of 7.7% was obtained [9]; relatively low compared to the work carried out at the hemodialysis center in the city of Fez which obtained a rate of 14.1% [12].
Nosocomial infections are responsible for a prolongation of the length of stay with an increase in the economic cost and are associated with a significant mortality. They are a real problem and go hand in hand with the evolution of medicine and care techniques. The microbial ecology of medical surfaces and devices is dense and consists of multi-resistant bacteria. Due to the high levels of contamination and the high threshold of positivity and resistance of all isolated bacteria affecting different units, it is important and urgent to evaluate and strengthen the disinfection practices of surfaces and medical devices at the Jourdan Medical Services. This study provides local and original data on the emergence of multi-drug resistant bacteria that could be responsible for the spread of nosocomial diseases. Limited resources, as well as local technical platform, did not allow us to perform molecular tests in order to type strains circulating in the hospital. The limited means did not allow us to carry out the sampling of all units present at the Jordan Medical Services.
What is known about this topic
- Micro-organisms involved in the hospital environment are most often multi-resistant to antibiotics, and the main source of diffusion of highly pathogenic strains in the departments.
What this study adds
- Indeed, this manuscript meets the objectives associated with the process of epidemiological surveillance of hospital infections, bringing added value to the control of antimicrobial resistance, a current scourge in our locality;
- The microbial ecology of medical surfaces and devices is dense and consists of multi-resistant bacteria;
- This study provides local and original data on the emergence of multi-drug resistant bacteria that could be responsible for the spread of nosocomial diseases.
The authors declare competing interests.
Nadege Lylian Doumou AMvomo designed the study and wrote the manuscript. Cybelle Kabit Sama Kemayou conducted the data collection. Nadege Lylian Doumou AMvomo, Cybelle Kabit Sama Kemayou, Bouopda Rodrigue and Esther Voundi Voundi carried out the statistical analysis. Bouopda Rodrigue, Esther Voundi Voundi, Flora Christelle Nkwinkwa Minkeza, Laure Patricia Mbopda, Cindy Loïda Yonga Tchoumi, Lesly Ornella Tchouangeum Kenmoe, Gilles Tchouassi Simeni, Eric Chokote Tolo, Ruth Ngongang critically read the manuscript. All authors read and approved the final version of the manuscript.
The authors would like to thank all the people who contributed to the realization of this work, especially the promoter of the Jourdan Medical Services and the heads of the units involved in this study.
Table 1: level of contamination per unit in our study
Table 2: distribution of isolated bacteria per unit in our study
Table 3: distribution of bacteria isolated by media in our study
Table 4: resistance profile of the isolated bacteria to the tested antibiotics
Figure 1: positivity threshold per unit in our study
- Baranovsky S. Circulation et persistance de pathogènes nosocomiaux multirésistants et hautement résistants émergents dans l´environnement hospitalier: complexité des unités de transmission (Circulation and persistence of emerging multidrug-resistant and highly resistant nosocomial pathogens in the hospital environment: complexity of transmission units). Université Montpellier. 2020. Google Scholar
- Ango PD, Konan KD, Kouamé KA, Sai SS, Tchimou AY, Adingra SC et al. Écologie Microbienne des Surfaces et Dispositifs Médicaux au Service de Réanimation du Centre Hospitalier et Universitaire (CHU) de Treichville (Microbial Ecology of Surfaces and Medical Devices in the Intensive Care Unit of the University Hospital Centre (CHU) of Treichville). Health Sci Dis. 2020;21(1). Google Scholar
- Ebongue CO, Nguatcheussi CM, Mefo´o JPN, Bollanga J, Adiogo D, Luma HN. Microbial Ecology and Antibiotic Susceptibility Profile of Germs Isolated from Hospital Surfaces and Medical Devices in a Reference Hospital in Douala (Cameroon). Adv Microbiol. 28 févr 2018;8(2):125-37. Google Scholar
- L´OMS. L´OMS publie le tout premier rapport mondial sur la lutte anti-infectieuse. Accessed Dec 13, 2022.
- Nabila S, Adil E, Abedelazi C, Nabila A, Samir H, Abdelmajid S. Rôle de l´environnement hospitalier dans la prévention des infections nosocomiales: Surveillance de la flore des surfaces à l´hôpital El Idrissi de Kenitra-Maroc. European Scientific Journal. 2014;10(9):237-47. Google Scholar
- ISO. ISO 14698-1:2003(en) Cleanrooms and associated controlled environments - Biocontamination control - Part 1: General principles and methods. Accessed Dec 13, 2022.
- 3M Sciences. guide-de-gestion-de-l'hygiene-relatif-aux-surfaces-environnementales. 2016. Accessed Dec 13, 2022.
- Société Française de Microbiologie. Comité de l´Antibiogramme de la SFM (CASFM) V1.0 Mai 2022. Accessed Dec 13, 2022.
- Chaoui L, Mhand R, Mellouki F, Rhallabi N. Contamination of the Surfaces of a Health Care Environment by Multidrug-Resistant (MDR) Bacteria. Int J Microbiol. 2019 Nov 29;2019:3236526. PubMed | Google Scholar
- Marty L, Jarlier V. Surveillance of multiresistant bacteria: justification, role of the laboratory, indicators, and recent French data. Pathol Biol (Paris). April 2017;46(4):217-26. PubMed | Google Scholar
- Berthelot P, Grattard F, Mallaval FO, Ros A, Lucht F, Pozzetto B. Épidémiologie des infections nosocomiales à Pseudomonas aeruginosa, Burkholderia cepacia et Stenotrophomonas maltophilia. Pathol Biol. 2005;53(6):341-8. Google Scholar
- Berrada S, Benjelloun Touimi G, Bennani L, Diarra AS, Oumokhtar B, El Ouali Lalami A et al. Exploration microbiologique des surfaces d´un centre d´hémodialyse de la ville de Fès: étude descriptive transversale. Rev Francoph Int Rech Infirm. Jun 1;3(2):120-8. Google Scholar